| Full name: proteasome 26S subunit, non-ATPase 13 | Alias Symbol: p40.5|Rpn9 | ||
| Type: protein-coding gene | Cytoband: 11p15.5 | ||
| Entrez ID: 5719 | HGNC ID: HGNC:9558 | Ensembl Gene: ENSG00000185627 | OMIM ID: 603481 |
| Drug and gene relationship at DGIdb | |||
Expression of PSMD13:
| Dataset | Gene | EntrezID | Probe | Log2FC | Adj.pValue | Expression |
|---|---|---|---|---|---|---|
| GSE17351 | PSMD13 | 5719 | 201232_s_at | 0.1831 | 0.6750 | |
| GSE20347 | PSMD13 | 5719 | 201232_s_at | 0.2579 | 0.0685 | |
| GSE23400 | PSMD13 | 5719 | 201232_s_at | 0.3089 | 0.0000 | |
| GSE26886 | PSMD13 | 5719 | 201232_s_at | -0.2431 | 0.1856 | |
| GSE29001 | PSMD13 | 5719 | 201232_s_at | 0.1509 | 0.6950 | |
| GSE38129 | PSMD13 | 5719 | 201232_s_at | 0.3848 | 0.0006 | |
| GSE45670 | PSMD13 | 5719 | 201232_s_at | 0.2857 | 0.0307 | |
| GSE53622 | PSMD13 | 5719 | 65156 | 0.2688 | 0.0000 | |
| GSE53624 | PSMD13 | 5719 | 65156 | 0.3696 | 0.0000 | |
| GSE63941 | PSMD13 | 5719 | 201232_s_at | -0.6670 | 0.1802 | |
| GSE77861 | PSMD13 | 5719 | 201232_s_at | 0.4306 | 0.0833 | |
| GSE97050 | PSMD13 | 5719 | A_23_P75889 | 0.4461 | 0.1403 | |
| SRP007169 | PSMD13 | 5719 | RNAseq | 2.2009 | 0.0009 | |
| SRP008496 | PSMD13 | 5719 | RNAseq | 2.5232 | 0.0000 | |
| SRP064894 | PSMD13 | 5719 | RNAseq | 0.3715 | 0.1042 | |
| SRP133303 | PSMD13 | 5719 | RNAseq | 0.3364 | 0.0079 | |
| SRP159526 | PSMD13 | 5719 | RNAseq | -0.0025 | 0.9885 | |
| SRP193095 | PSMD13 | 5719 | RNAseq | 0.1268 | 0.2428 | |
| SRP219564 | PSMD13 | 5719 | RNAseq | 0.2143 | 0.5056 | |
| TCGA | PSMD13 | 5719 | RNAseq | 0.1123 | 0.0168 |
Upregulated datasets: 2; Downregulated datasets: 0.
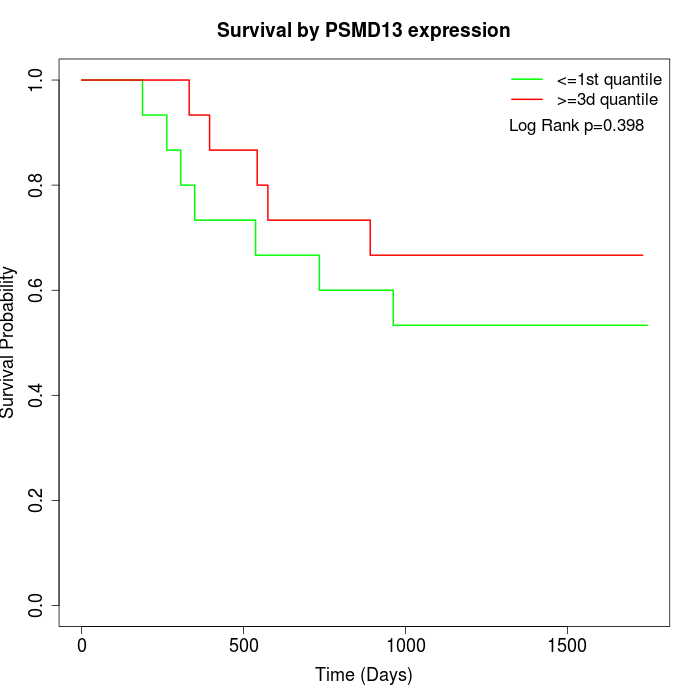
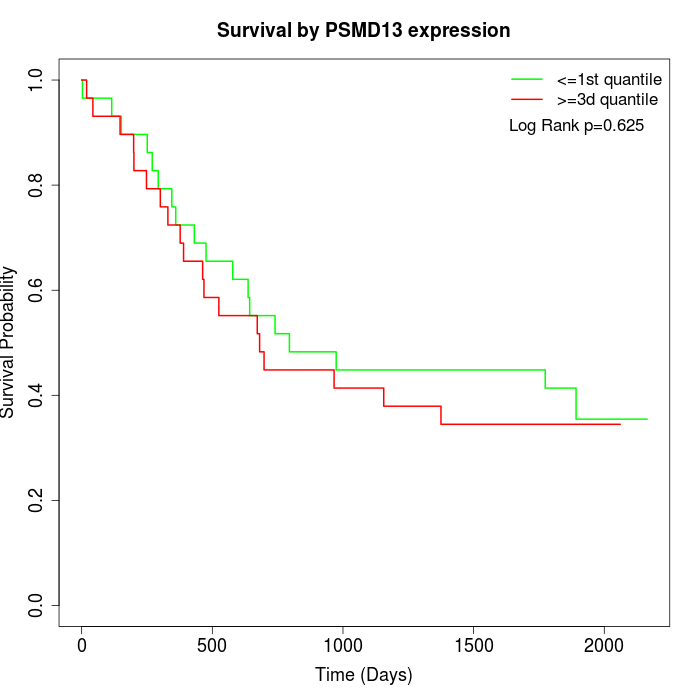
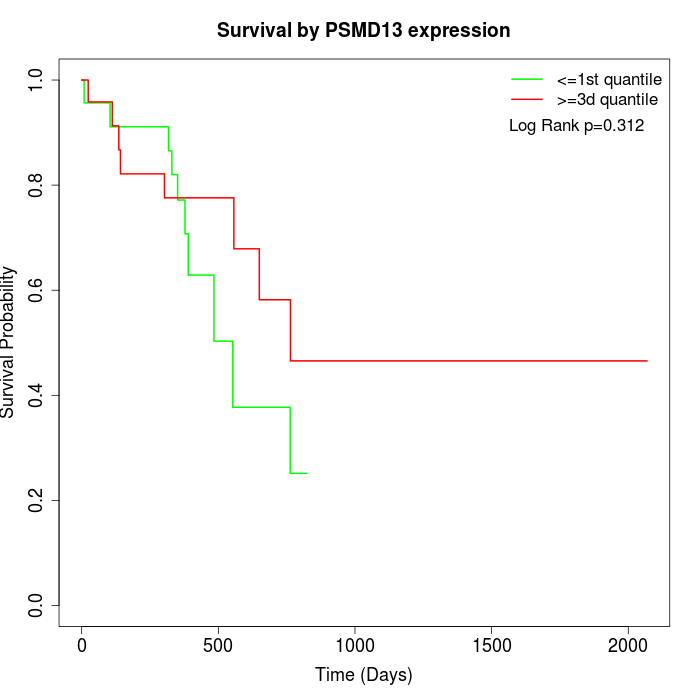
Survival by PSMD13 expression:
Note: Click image to view full size file.
Copy number change of PSMD13:
| Dataset | Gene | EntrezID | Gain | Loss | Normal | Detail |
|---|---|---|---|---|---|---|
| GSE15526 | PSMD13 | 5719 | 1 | 11 | 18 | |
| GSE20123 | PSMD13 | 5719 | 1 | 12 | 17 | |
| GSE43470 | PSMD13 | 5719 | 1 | 11 | 31 | |
| GSE46452 | PSMD13 | 5719 | 7 | 8 | 44 | |
| GSE47630 | PSMD13 | 5719 | 4 | 12 | 24 | |
| GSE54993 | PSMD13 | 5719 | 3 | 2 | 65 | |
| GSE54994 | PSMD13 | 5719 | 1 | 12 | 40 | |
| GSE60625 | PSMD13 | 5719 | 0 | 0 | 11 | |
| GSE74703 | PSMD13 | 5719 | 1 | 9 | 26 | |
| GSE74704 | PSMD13 | 5719 | 0 | 8 | 12 | |
| TCGA | PSMD13 | 5719 | 8 | 38 | 50 |
Total number of gains: 27; Total number of losses: 123; Total Number of normals: 338.
Somatic mutations of PSMD13:
Generating mutation plots.
Highly correlated genes for PSMD13:
Showing top 20/749 corelated genes with mean PCC>0.5.
| Gene1 | Gene2 | Mean PCC | Num. Datasets | Num. PCC<0 | Num. PCC>0.5 |
|---|---|---|---|---|---|
| PSMD13 | TRAF7 | 0.740229 | 4 | 0 | 4 |
| PSMD13 | DUS3L | 0.708658 | 3 | 0 | 3 |
| PSMD13 | PRMT6 | 0.705463 | 3 | 0 | 3 |
| PSMD13 | ARL13B | 0.703168 | 3 | 0 | 3 |
| PSMD13 | LY96 | 0.695059 | 3 | 0 | 3 |
| PSMD13 | SPRED2 | 0.693631 | 4 | 0 | 4 |
| PSMD13 | IFNAR2 | 0.689917 | 3 | 0 | 3 |
| PSMD13 | NECAP1 | 0.68637 | 3 | 0 | 3 |
| PSMD13 | PSMG3 | 0.686141 | 6 | 0 | 5 |
| PSMD13 | SASS6 | 0.685967 | 4 | 0 | 3 |
| PSMD13 | SLC37A3 | 0.683688 | 3 | 0 | 3 |
| PSMD13 | ID1 | 0.678926 | 3 | 0 | 3 |
| PSMD13 | SPSB2 | 0.678298 | 3 | 0 | 3 |
| PSMD13 | ABCB6 | 0.676159 | 4 | 0 | 3 |
| PSMD13 | SNAP23 | 0.671895 | 3 | 0 | 3 |
| PSMD13 | RAD18 | 0.66964 | 3 | 0 | 3 |
| PSMD13 | INTS9 | 0.669295 | 4 | 0 | 4 |
| PSMD13 | SAAL1 | 0.669294 | 6 | 0 | 5 |
| PSMD13 | STX10 | 0.668767 | 4 | 0 | 4 |
| PSMD13 | MRPL37 | 0.668654 | 5 | 0 | 4 |
For details and further investigation, click here