| Full name: RNA binding motif protein 25 | Alias Symbol: S164|fSAP94|NET52|Snu71 | ||
| Type: protein-coding gene | Cytoband: 14q24.2 | ||
| Entrez ID: 58517 | HGNC ID: HGNC:23244 | Ensembl Gene: ENSG00000119707 | OMIM ID: 612427 |
Screen Evidence:
| |||
Expression of RBM25:
| Dataset | Gene | EntrezID | Probe | Log2FC | Adj.pValue | Expression |
|---|---|---|---|---|---|---|
| GSE17351 | RBM25 | 58517 | 212033_at | 0.2500 | 0.6401 | |
| GSE20347 | RBM25 | 58517 | 212033_at | 0.1492 | 0.2819 | |
| GSE23400 | RBM25 | 58517 | 212033_at | 0.2439 | 0.0009 | |
| GSE26886 | RBM25 | 58517 | 212027_at | 0.8402 | 0.0310 | |
| GSE29001 | RBM25 | 58517 | 212031_at | 0.2303 | 0.3479 | |
| GSE38129 | RBM25 | 58517 | 212033_at | 0.0987 | 0.3648 | |
| GSE45670 | RBM25 | 58517 | 212033_at | -0.0366 | 0.8053 | |
| GSE53622 | RBM25 | 58517 | 92800 | 0.1515 | 0.0021 | |
| GSE53624 | RBM25 | 58517 | 92800 | 0.0377 | 0.4351 | |
| GSE63941 | RBM25 | 58517 | 212033_at | -0.7099 | 0.0996 | |
| GSE77861 | RBM25 | 58517 | 212033_at | -0.0903 | 0.7964 | |
| GSE97050 | RBM25 | 58517 | A_23_P99762 | -0.0076 | 0.9783 | |
| SRP007169 | RBM25 | 58517 | RNAseq | 0.7124 | 0.1022 | |
| SRP008496 | RBM25 | 58517 | RNAseq | 0.7491 | 0.0095 | |
| SRP064894 | RBM25 | 58517 | RNAseq | -0.0238 | 0.9058 | |
| SRP133303 | RBM25 | 58517 | RNAseq | 0.0389 | 0.7385 | |
| SRP159526 | RBM25 | 58517 | RNAseq | -0.2194 | 0.3950 | |
| SRP193095 | RBM25 | 58517 | RNAseq | 0.0265 | 0.7971 | |
| SRP219564 | RBM25 | 58517 | RNAseq | 0.0461 | 0.8755 | |
| TCGA | RBM25 | 58517 | RNAseq | -0.0475 | 0.2461 |
Upregulated datasets: 0; Downregulated datasets: 0.
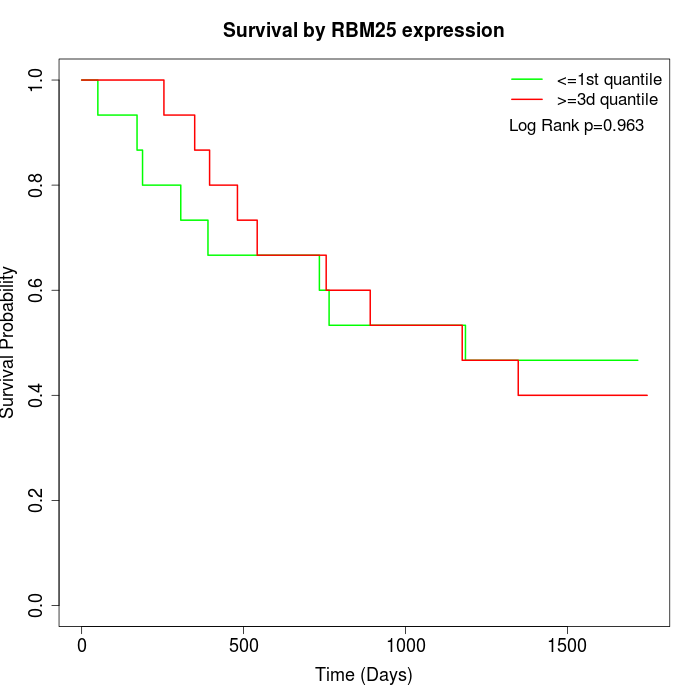
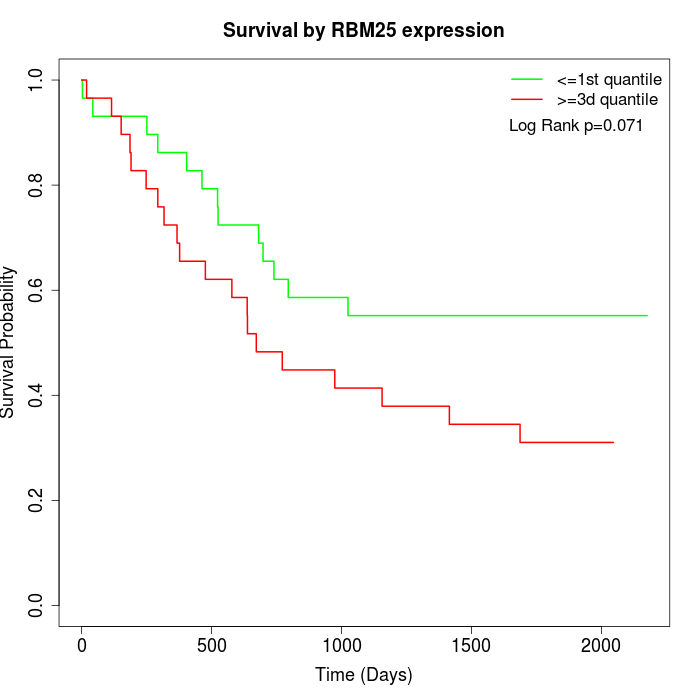
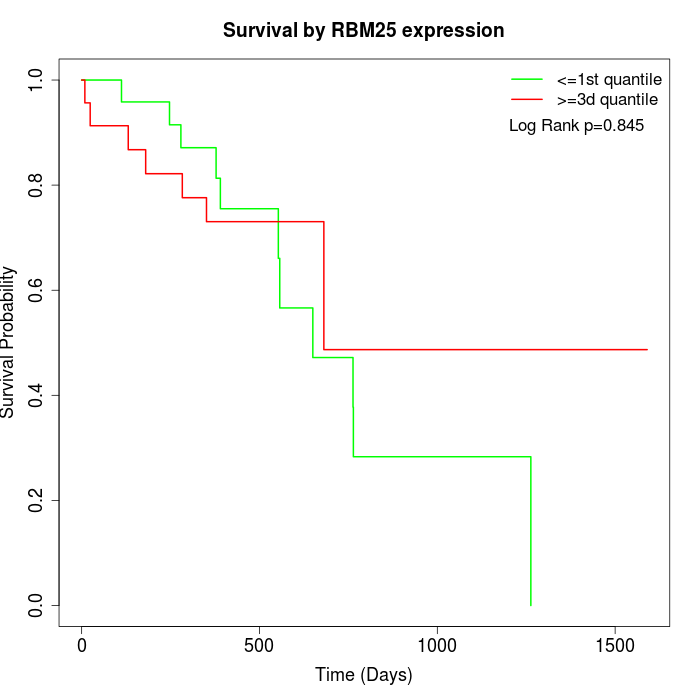
Survival by RBM25 expression:
Note: Click image to view full size file.
Copy number change of RBM25:
| Dataset | Gene | EntrezID | Gain | Loss | Normal | Detail |
|---|---|---|---|---|---|---|
| GSE15526 | RBM25 | 58517 | 8 | 2 | 20 | |
| GSE20123 | RBM25 | 58517 | 8 | 2 | 20 | |
| GSE43470 | RBM25 | 58517 | 8 | 3 | 32 | |
| GSE46452 | RBM25 | 58517 | 16 | 3 | 40 | |
| GSE47630 | RBM25 | 58517 | 11 | 8 | 21 | |
| GSE54993 | RBM25 | 58517 | 3 | 8 | 59 | |
| GSE54994 | RBM25 | 58517 | 19 | 4 | 30 | |
| GSE60625 | RBM25 | 58517 | 0 | 2 | 9 | |
| GSE74703 | RBM25 | 58517 | 7 | 3 | 26 | |
| GSE74704 | RBM25 | 58517 | 3 | 2 | 15 | |
| TCGA | RBM25 | 58517 | 31 | 17 | 48 |
Total number of gains: 114; Total number of losses: 54; Total Number of normals: 320.
Somatic mutations of RBM25:
Generating mutation plots.
Highly correlated genes for RBM25:
Showing top 20/64 corelated genes with mean PCC>0.5.
| Gene1 | Gene2 | Mean PCC | Num. Datasets | Num. PCC<0 | Num. PCC>0.5 |
|---|---|---|---|---|---|
| RBM25 | DNAJC16 | 0.747914 | 3 | 0 | 3 |
| RBM25 | ZNF678 | 0.723992 | 3 | 0 | 3 |
| RBM25 | PARS2 | 0.680683 | 3 | 0 | 3 |
| RBM25 | MDC1 | 0.670908 | 3 | 0 | 3 |
| RBM25 | C19orf25 | 0.670652 | 3 | 0 | 3 |
| RBM25 | ANKS3 | 0.669199 | 3 | 0 | 3 |
| RBM25 | RASL11B | 0.659909 | 4 | 0 | 4 |
| RBM25 | PEX26 | 0.655869 | 3 | 0 | 3 |
| RBM25 | GPC2 | 0.652902 | 3 | 0 | 3 |
| RBM25 | CDC42BPB | 0.652507 | 4 | 0 | 4 |
| RBM25 | ATG4D | 0.642452 | 3 | 0 | 3 |
| RBM25 | RAB40B | 0.625847 | 4 | 0 | 3 |
| RBM25 | CLCN6 | 0.624776 | 5 | 0 | 4 |
| RBM25 | DOHH | 0.622143 | 4 | 0 | 3 |
| RBM25 | BCAT2 | 0.621144 | 4 | 0 | 3 |
| RBM25 | AP1M1 | 0.620755 | 4 | 0 | 3 |
| RBM25 | ZNF205 | 0.617131 | 4 | 0 | 4 |
| RBM25 | SHARPIN | 0.616026 | 3 | 0 | 3 |
| RBM25 | ATPAF1 | 0.612182 | 3 | 0 | 3 |
| RBM25 | POMT2 | 0.609585 | 5 | 0 | 3 |
For details and further investigation, click here