| Full name: splicing factor SWAP | Alias Symbol: SWAP | ||
| Type: protein-coding gene | Cytoband: 12q24.33 | ||
| Entrez ID: 6433 | HGNC ID: HGNC:10790 | Ensembl Gene: ENSG00000061936 | OMIM ID: 601945 |
| Drug and gene relationship at DGIdb | |||
Screen Evidence:
| |||
Expression of SFSWAP:
| Dataset | Gene | EntrezID | Probe | Log2FC | Adj.pValue | Expression |
|---|---|---|---|---|---|---|
| GSE17351 | SFSWAP | 6433 | 202774_s_at | 0.3224 | 0.6676 | |
| GSE20347 | SFSWAP | 6433 | 202774_s_at | -0.0566 | 0.7555 | |
| GSE23400 | SFSWAP | 6433 | 202774_s_at | 0.0026 | 0.9594 | |
| GSE26886 | SFSWAP | 6433 | 202774_s_at | 0.3008 | 0.1043 | |
| GSE29001 | SFSWAP | 6433 | 202774_s_at | 0.0143 | 0.9672 | |
| GSE38129 | SFSWAP | 6433 | 202774_s_at | -0.1039 | 0.3853 | |
| GSE45670 | SFSWAP | 6433 | 202774_s_at | -0.0732 | 0.5493 | |
| GSE53622 | SFSWAP | 6433 | 57655 | 0.2396 | 0.0000 | |
| GSE53624 | SFSWAP | 6433 | 57655 | 0.2758 | 0.0000 | |
| GSE63941 | SFSWAP | 6433 | 202774_s_at | 0.0500 | 0.8410 | |
| GSE77861 | SFSWAP | 6433 | 202774_s_at | 0.2589 | 0.1076 | |
| GSE97050 | SFSWAP | 6433 | A_32_P109704 | 0.0037 | 0.9896 | |
| SRP007169 | SFSWAP | 6433 | RNAseq | 0.2654 | 0.5708 | |
| SRP008496 | SFSWAP | 6433 | RNAseq | 0.1616 | 0.5893 | |
| SRP064894 | SFSWAP | 6433 | RNAseq | 0.0614 | 0.5790 | |
| SRP133303 | SFSWAP | 6433 | RNAseq | -0.0373 | 0.7011 | |
| SRP159526 | SFSWAP | 6433 | RNAseq | 0.1289 | 0.6186 | |
| SRP193095 | SFSWAP | 6433 | RNAseq | 0.3069 | 0.0168 | |
| SRP219564 | SFSWAP | 6433 | RNAseq | 0.2151 | 0.5549 |
Upregulated datasets: 0; Downregulated datasets: 0.
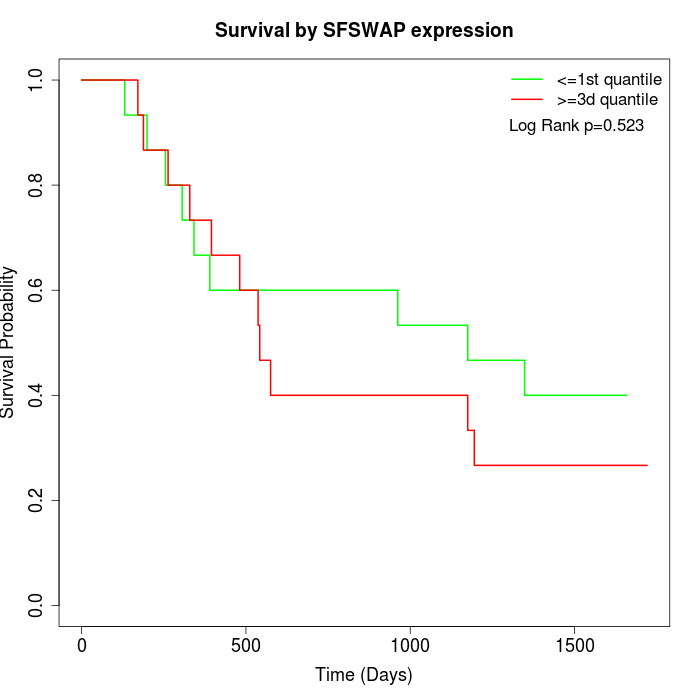
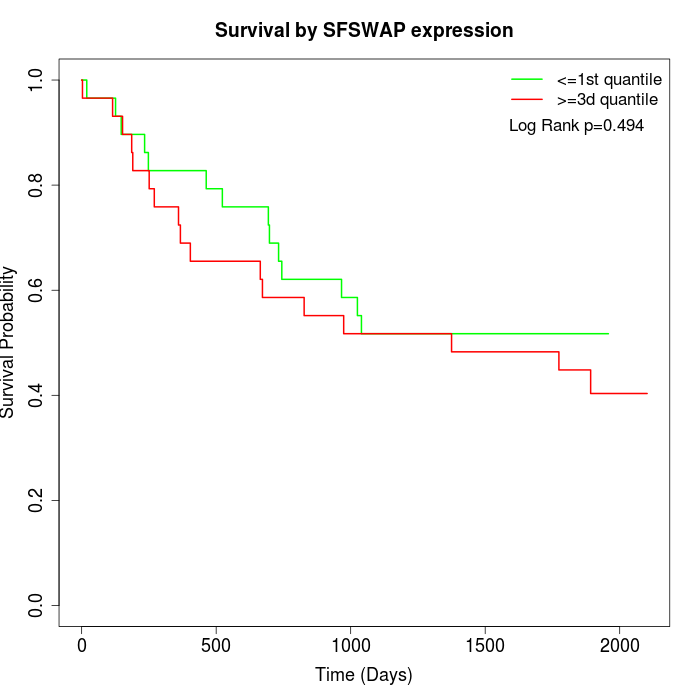
Survival by SFSWAP expression:
Note: Click image to view full size file.
Copy number change of SFSWAP:
| Dataset | Gene | EntrezID | Gain | Loss | Normal | Detail |
|---|---|---|---|---|---|---|
| GSE15526 | SFSWAP | 6433 | 4 | 5 | 21 | |
| GSE20123 | SFSWAP | 6433 | 4 | 5 | 21 | |
| GSE43470 | SFSWAP | 6433 | 3 | 3 | 37 | |
| GSE46452 | SFSWAP | 6433 | 8 | 3 | 48 | |
| GSE47630 | SFSWAP | 6433 | 9 | 3 | 28 | |
| GSE54993 | SFSWAP | 6433 | 0 | 6 | 64 | |
| GSE54994 | SFSWAP | 6433 | 4 | 8 | 41 | |
| GSE60625 | SFSWAP | 6433 | 0 | 0 | 11 | |
| GSE74703 | SFSWAP | 6433 | 2 | 3 | 31 | |
| GSE74704 | SFSWAP | 6433 | 2 | 3 | 15 | |
| TCGA | SFSWAP | 6433 | 22 | 13 | 61 |
Total number of gains: 58; Total number of losses: 52; Total Number of normals: 378.
Somatic mutations of SFSWAP:
Generating mutation plots.
Highly correlated genes for SFSWAP:
Showing top 20/364 corelated genes with mean PCC>0.5.
| Gene1 | Gene2 | Mean PCC | Num. Datasets | Num. PCC<0 | Num. PCC>0.5 |
|---|---|---|---|---|---|
| SFSWAP | TIPRL | 0.854189 | 3 | 0 | 3 |
| SFSWAP | ZMYND19 | 0.811452 | 3 | 0 | 3 |
| SFSWAP | SURF6 | 0.795856 | 3 | 0 | 3 |
| SFSWAP | IRF2BP2 | 0.794649 | 3 | 0 | 3 |
| SFSWAP | NPHP3 | 0.781176 | 3 | 0 | 3 |
| SFSWAP | PPP1R16A | 0.779918 | 3 | 0 | 3 |
| SFSWAP | TMEM181 | 0.778206 | 3 | 0 | 3 |
| SFSWAP | NOL7 | 0.774024 | 3 | 0 | 3 |
| SFSWAP | FIP1L1 | 0.772289 | 4 | 0 | 4 |
| SFSWAP | ZNF121 | 0.764515 | 3 | 0 | 3 |
| SFSWAP | ZBTB2 | 0.758175 | 3 | 0 | 3 |
| SFSWAP | POLR3C | 0.754781 | 3 | 0 | 3 |
| SFSWAP | ZNF766 | 0.754673 | 3 | 0 | 3 |
| SFSWAP | TERF1 | 0.75066 | 4 | 0 | 4 |
| SFSWAP | HIRA | 0.750356 | 3 | 0 | 3 |
| SFSWAP | RBM26 | 0.746828 | 4 | 0 | 4 |
| SFSWAP | ZNF124 | 0.746103 | 3 | 0 | 3 |
| SFSWAP | GAR1 | 0.744814 | 3 | 0 | 3 |
| SFSWAP | CCT8 | 0.743428 | 3 | 0 | 3 |
| SFSWAP | MRPS6 | 0.741072 | 3 | 0 | 3 |
For details and further investigation, click here