| Full name: solute carrier family 12 member 2 | Alias Symbol: NKCC1|BSC|BSC2|PPP1R141 | ||
| Type: protein-coding gene | Cytoband: 5q23.3 | ||
| Entrez ID: 6558 | HGNC ID: HGNC:10911 | Ensembl Gene: ENSG00000064651 | OMIM ID: 600840 |
| Related drugs: ALDOSTERONE, BUMETANIDE, FUROSEMIDE, PIRETANIDE... [more] | |||
SLC12A2 involved pathways:
| KEGG pathway | Description | View |
|---|---|---|
| hsa05110 | Vibrio cholerae infection |
Expression of SLC12A2:
| Dataset | Gene | EntrezID | Probe | Log2FC | Adj.pValue | Expression |
|---|---|---|---|---|---|---|
| GSE17351 | SLC12A2 | 6558 | 225835_at | -0.5546 | 0.5873 | |
| GSE20347 | SLC12A2 | 6558 | 204404_at | 0.6861 | 0.0067 | |
| GSE23400 | SLC12A2 | 6558 | 204404_at | 0.0380 | 0.4253 | |
| GSE26886 | SLC12A2 | 6558 | 225835_at | 0.5944 | 0.1083 | |
| GSE29001 | SLC12A2 | 6558 | 204404_at | 0.6797 | 0.0306 | |
| GSE38129 | SLC12A2 | 6558 | 204404_at | 0.4429 | 0.0871 | |
| GSE45670 | SLC12A2 | 6558 | 225835_at | -0.3148 | 0.4353 | |
| GSE53622 | SLC12A2 | 6558 | 18518 | -0.1838 | 0.3935 | |
| GSE53624 | SLC12A2 | 6558 | 50715 | -0.2570 | 0.0790 | |
| GSE63941 | SLC12A2 | 6558 | 225835_at | -0.2936 | 0.7232 | |
| GSE77861 | SLC12A2 | 6558 | 204404_at | 0.4548 | 0.1069 | |
| GSE97050 | SLC12A2 | 6558 | A_32_P25437 | -0.6117 | 0.2857 | |
| SRP007169 | SLC12A2 | 6558 | RNAseq | 1.1920 | 0.0058 | |
| SRP008496 | SLC12A2 | 6558 | RNAseq | 1.0108 | 0.0001 | |
| SRP064894 | SLC12A2 | 6558 | RNAseq | -0.3761 | 0.2573 | |
| SRP133303 | SLC12A2 | 6558 | RNAseq | -0.4942 | 0.2145 | |
| SRP159526 | SLC12A2 | 6558 | RNAseq | 0.9996 | 0.0301 | |
| SRP193095 | SLC12A2 | 6558 | RNAseq | -0.5904 | 0.1529 | |
| SRP219564 | SLC12A2 | 6558 | RNAseq | 0.0769 | 0.9255 | |
| TCGA | SLC12A2 | 6558 | RNAseq | -0.6747 | 0.0000 |
Upregulated datasets: 2; Downregulated datasets: 0.
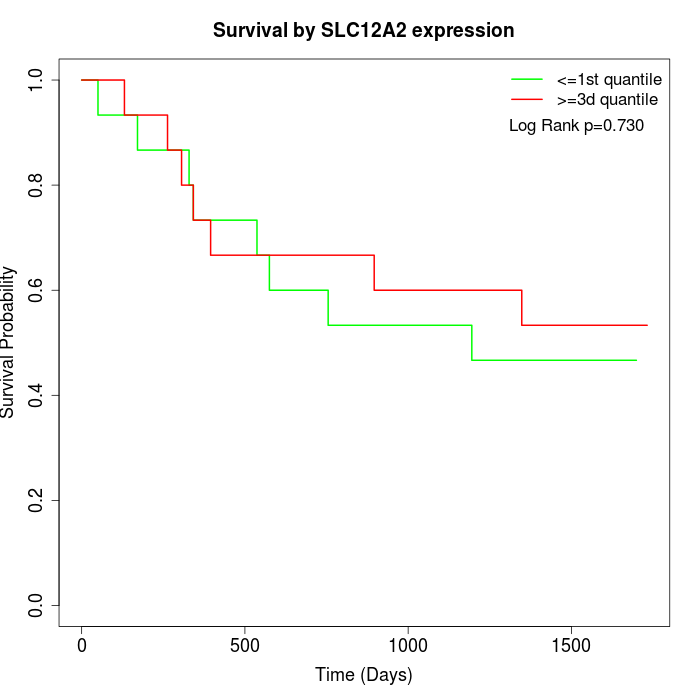
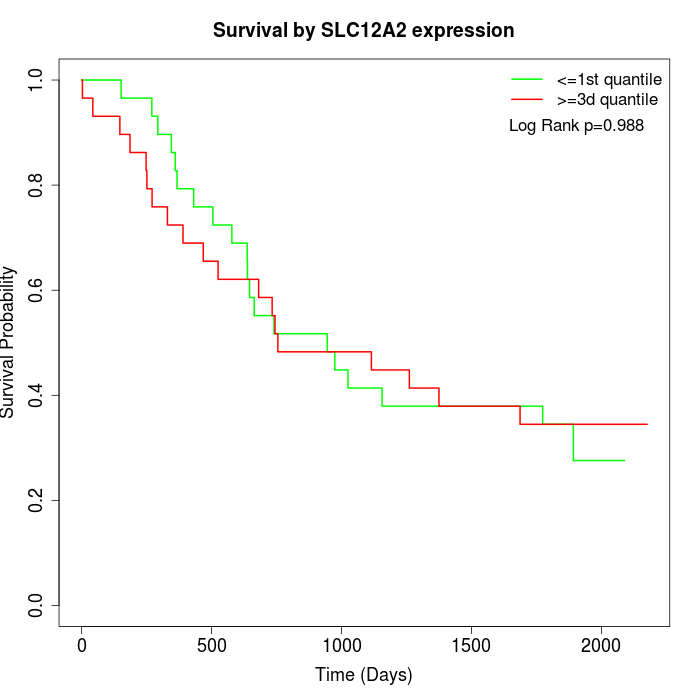
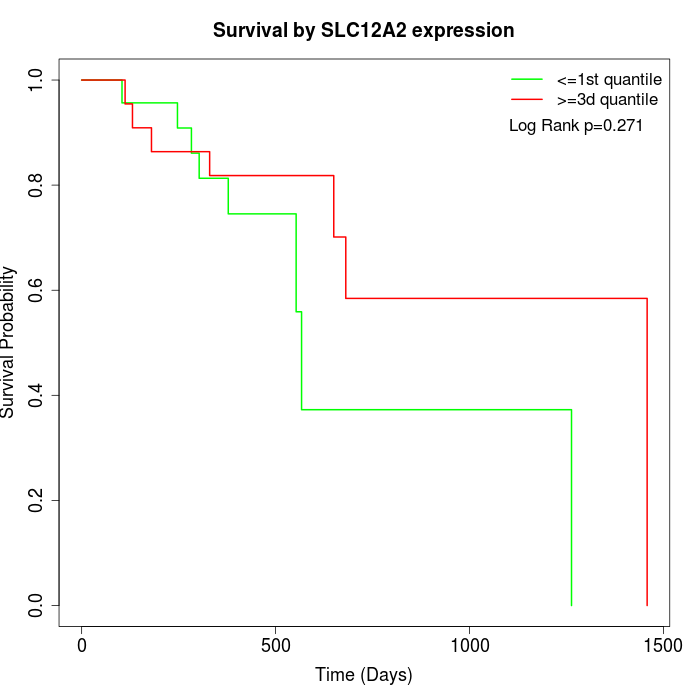
Survival by SLC12A2 expression:
Note: Click image to view full size file.
Copy number change of SLC12A2:
| Dataset | Gene | EntrezID | Gain | Loss | Normal | Detail |
|---|---|---|---|---|---|---|
| GSE15526 | SLC12A2 | 6558 | 1 | 12 | 17 | |
| GSE20123 | SLC12A2 | 6558 | 1 | 12 | 17 | |
| GSE43470 | SLC12A2 | 6558 | 3 | 8 | 32 | |
| GSE46452 | SLC12A2 | 6558 | 0 | 27 | 32 | |
| GSE47630 | SLC12A2 | 6558 | 0 | 21 | 19 | |
| GSE54993 | SLC12A2 | 6558 | 9 | 1 | 60 | |
| GSE54994 | SLC12A2 | 6558 | 2 | 14 | 37 | |
| GSE60625 | SLC12A2 | 6558 | 0 | 0 | 11 | |
| GSE74703 | SLC12A2 | 6558 | 2 | 6 | 28 | |
| GSE74704 | SLC12A2 | 6558 | 1 | 6 | 13 | |
| TCGA | SLC12A2 | 6558 | 2 | 40 | 54 |
Total number of gains: 21; Total number of losses: 147; Total Number of normals: 320.
Somatic mutations of SLC12A2:
Generating mutation plots.
Highly correlated genes for SLC12A2:
Showing top 20/180 corelated genes with mean PCC>0.5.
| Gene1 | Gene2 | Mean PCC | Num. Datasets | Num. PCC<0 | Num. PCC>0.5 |
|---|---|---|---|---|---|
| SLC12A2 | TPPP3 | 0.771359 | 3 | 0 | 3 |
| SLC12A2 | ANKS1A | 0.688701 | 3 | 0 | 3 |
| SLC12A2 | CAP2 | 0.675822 | 3 | 0 | 3 |
| SLC12A2 | UST | 0.675656 | 3 | 0 | 3 |
| SLC12A2 | CC2D2A | 0.672201 | 4 | 0 | 3 |
| SLC12A2 | IL13RA2 | 0.669092 | 3 | 0 | 3 |
| SLC12A2 | SPRY4 | 0.66101 | 4 | 0 | 3 |
| SLC12A2 | NPTX1 | 0.653613 | 3 | 0 | 3 |
| SLC12A2 | LAMA2 | 0.643941 | 4 | 0 | 3 |
| SLC12A2 | KITLG | 0.64145 | 3 | 0 | 3 |
| SLC12A2 | RANGRF | 0.635256 | 3 | 0 | 3 |
| SLC12A2 | PYGB | 0.634788 | 5 | 0 | 5 |
| SLC12A2 | SLC2A12 | 0.625281 | 3 | 0 | 3 |
| SLC12A2 | ENC1 | 0.62182 | 3 | 0 | 3 |
| SLC12A2 | DCAF17 | 0.621194 | 4 | 0 | 3 |
| SLC12A2 | SSRP1 | 0.616458 | 3 | 0 | 3 |
| SLC12A2 | DDHD2 | 0.614895 | 5 | 0 | 3 |
| SLC12A2 | TSC22D1 | 0.612996 | 4 | 0 | 4 |
| SLC12A2 | APAF1 | 0.610725 | 3 | 0 | 3 |
| SLC12A2 | DYNC2H1 | 0.60923 | 4 | 0 | 3 |
For details and further investigation, click here