| Full name: APC membrane recruitment protein 2 | Alias Symbol: FLJ25477 | ||
| Type: protein-coding gene | Cytoband: 13q12.13 | ||
| Entrez ID: 219287 | HGNC ID: HGNC:26360 | Ensembl Gene: ENSG00000165566 | OMIM ID: 614659 |
Expression of AMER2:
| Dataset | Gene | EntrezID | Probe | Log2FC | Adj.pValue | Expression |
|---|---|---|---|---|---|---|
| GSE17351 | AMER2 | 219287 | 1553720_a_at | -0.3129 | 0.1535 | |
| GSE26886 | AMER2 | 219287 | 1553720_a_at | -0.3614 | 0.0082 | |
| GSE45670 | AMER2 | 219287 | 1553720_a_at | 0.0375 | 0.7982 | |
| GSE53622 | AMER2 | 219287 | 28389 | 0.0871 | 0.5788 | |
| GSE53624 | AMER2 | 219287 | 28389 | 0.6715 | 0.0000 | |
| GSE63941 | AMER2 | 219287 | 1553720_a_at | 0.1446 | 0.4426 | |
| GSE77861 | AMER2 | 219287 | 1553720_a_at | -0.3849 | 0.0057 | |
| SRP133303 | AMER2 | 219287 | RNAseq | 0.1126 | 0.5410 | |
| SRP193095 | AMER2 | 219287 | RNAseq | -0.2068 | 0.2575 | |
| SRP219564 | AMER2 | 219287 | RNAseq | 0.4377 | 0.3060 |
Upregulated datasets: 0; Downregulated datasets: 0.
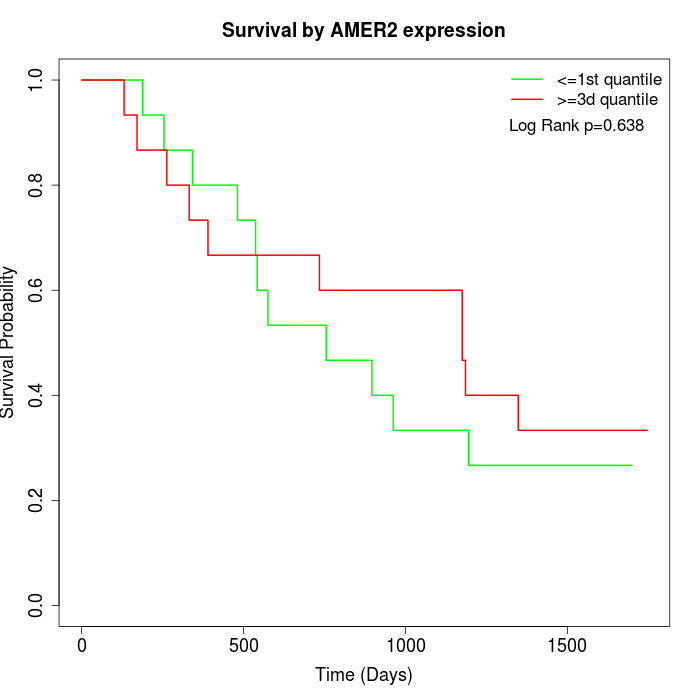
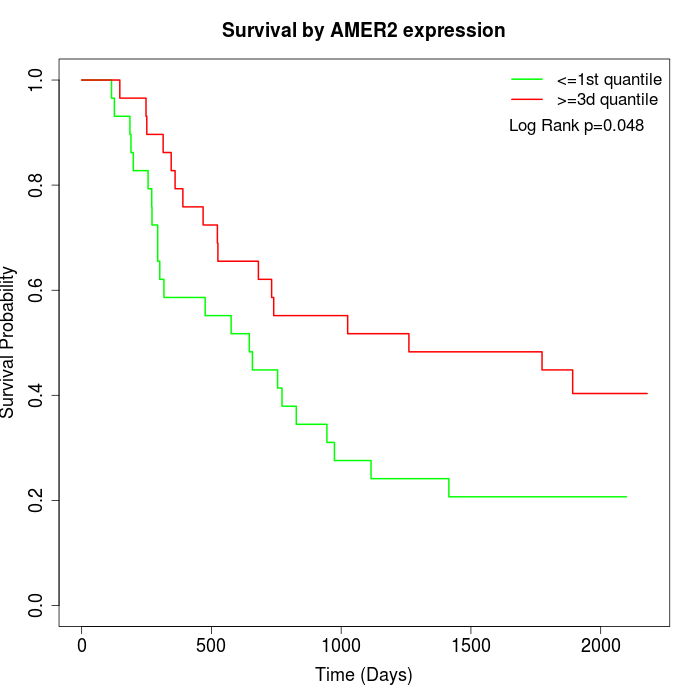
Survival by AMER2 expression:
Note: Click image to view full size file.
Copy number change of AMER2:
| Dataset | Gene | EntrezID | Gain | Loss | Normal | Detail |
|---|---|---|---|---|---|---|
| GSE15526 | AMER2 | 219287 | 1 | 12 | 17 | |
| GSE20123 | AMER2 | 219287 | 1 | 11 | 18 | |
| GSE43470 | AMER2 | 219287 | 2 | 12 | 29 | |
| GSE46452 | AMER2 | 219287 | 0 | 33 | 26 | |
| GSE47630 | AMER2 | 219287 | 3 | 26 | 11 | |
| GSE54993 | AMER2 | 219287 | 11 | 2 | 57 | |
| GSE54994 | AMER2 | 219287 | 1 | 13 | 39 | |
| GSE60625 | AMER2 | 219287 | 0 | 3 | 8 | |
| GSE74703 | AMER2 | 219287 | 2 | 9 | 25 | |
| GSE74704 | AMER2 | 219287 | 0 | 9 | 11 | |
| TCGA | AMER2 | 219287 | 5 | 46 | 45 |
Total number of gains: 26; Total number of losses: 176; Total Number of normals: 286.
Somatic mutations of AMER2:
Generating mutation plots.
Highly correlated genes for AMER2:
Showing top 20/286 corelated genes with mean PCC>0.5.
| Gene1 | Gene2 | Mean PCC | Num. Datasets | Num. PCC<0 | Num. PCC>0.5 |
|---|---|---|---|---|---|
| AMER2 | KRT33A | 0.780489 | 3 | 0 | 3 |
| AMER2 | LMAN2 | 0.744326 | 3 | 0 | 3 |
| AMER2 | BBOX1 | 0.740747 | 3 | 0 | 3 |
| AMER2 | MUC17 | 0.733691 | 3 | 0 | 3 |
| AMER2 | SMPD2 | 0.732816 | 3 | 0 | 3 |
| AMER2 | TP53I3 | 0.730149 | 3 | 0 | 3 |
| AMER2 | RNH1 | 0.726083 | 3 | 0 | 3 |
| AMER2 | HTR3B | 0.723315 | 3 | 0 | 3 |
| AMER2 | QSOX1 | 0.71934 | 3 | 0 | 3 |
| AMER2 | LINC00315 | 0.708969 | 3 | 0 | 3 |
| AMER2 | CNPPD1 | 0.704626 | 3 | 0 | 3 |
| AMER2 | GLI4 | 0.701454 | 4 | 0 | 4 |
| AMER2 | GALR3 | 0.700205 | 3 | 0 | 3 |
| AMER2 | CRABP2 | 0.699475 | 4 | 0 | 3 |
| AMER2 | CYP3A43 | 0.698214 | 3 | 0 | 3 |
| AMER2 | GAB2 | 0.698158 | 3 | 0 | 3 |
| AMER2 | SH3GL1 | 0.697811 | 3 | 0 | 3 |
| AMER2 | PHYHD1 | 0.69778 | 3 | 0 | 3 |
| AMER2 | PRDX5 | 0.694972 | 3 | 0 | 3 |
| AMER2 | CXCR2 | 0.692834 | 4 | 0 | 3 |
For details and further investigation, click here