| Full name: ATPase Na+/K+ transporting subunit alpha 1 | Alias Symbol: | ||
| Type: protein-coding gene | Cytoband: 1p13.1 | ||
| Entrez ID: 476 | HGNC ID: HGNC:799 | Ensembl Gene: ENSG00000163399 | OMIM ID: 182310 |
| Related drugs: ACETYLDIGITOXIN, ALMITRINE, BEPRIDIL, CHEMBL38827, CHEMBL444449, CICLOPIROX, DESLANOSIDE, DIGITOXIN, DIGOXIN, ISTAROXIME... [more] | |||
Screen Evidence:
| |||
ATP1A1 involved pathways:
Expression of ATP1A1:
| Dataset | Gene | EntrezID | Probe | Log2FC | Adj.pValue | Expression |
|---|---|---|---|---|---|---|
| GSE17351 | ATP1A1 | 476 | 220948_s_at | 0.2646 | 0.5751 | |
| GSE20347 | ATP1A1 | 476 | 220948_s_at | 0.2149 | 0.3249 | |
| GSE23400 | ATP1A1 | 476 | 220948_s_at | 0.5895 | 0.0000 | |
| GSE26886 | ATP1A1 | 476 | 220948_s_at | -0.1327 | 0.6736 | |
| GSE29001 | ATP1A1 | 476 | 220948_s_at | 0.4124 | 0.3453 | |
| GSE38129 | ATP1A1 | 476 | 220948_s_at | 0.5284 | 0.0010 | |
| GSE45670 | ATP1A1 | 476 | 220948_s_at | 0.5753 | 0.0024 | |
| GSE53622 | ATP1A1 | 476 | 85590 | 0.6845 | 0.0000 | |
| GSE53624 | ATP1A1 | 476 | 85590 | 0.8128 | 0.0000 | |
| GSE63941 | ATP1A1 | 476 | 220948_s_at | 0.2743 | 0.5765 | |
| GSE77861 | ATP1A1 | 476 | 220948_s_at | 0.1663 | 0.7150 | |
| GSE97050 | ATP1A1 | 476 | A_23_P1072 | 0.5972 | 0.1496 | |
| SRP007169 | ATP1A1 | 476 | RNAseq | 0.3842 | 0.3078 | |
| SRP008496 | ATP1A1 | 476 | RNAseq | 0.2548 | 0.2915 | |
| SRP064894 | ATP1A1 | 476 | RNAseq | 0.2586 | 0.1831 | |
| SRP133303 | ATP1A1 | 476 | RNAseq | 0.4939 | 0.0194 | |
| SRP159526 | ATP1A1 | 476 | RNAseq | 0.0734 | 0.7758 | |
| SRP193095 | ATP1A1 | 476 | RNAseq | 0.0958 | 0.5808 | |
| SRP219564 | ATP1A1 | 476 | RNAseq | 0.5938 | 0.2457 | |
| TCGA | ATP1A1 | 476 | RNAseq | 0.0439 | 0.3612 |
Upregulated datasets: 0; Downregulated datasets: 0.
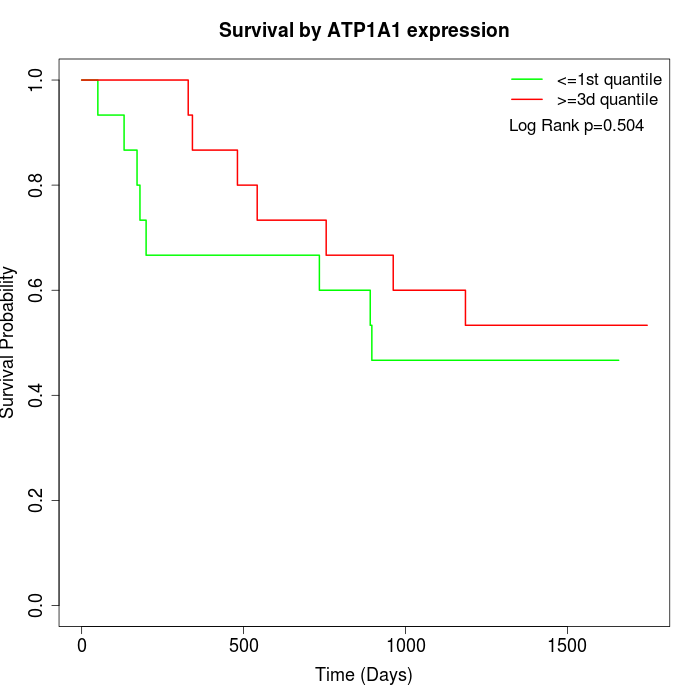
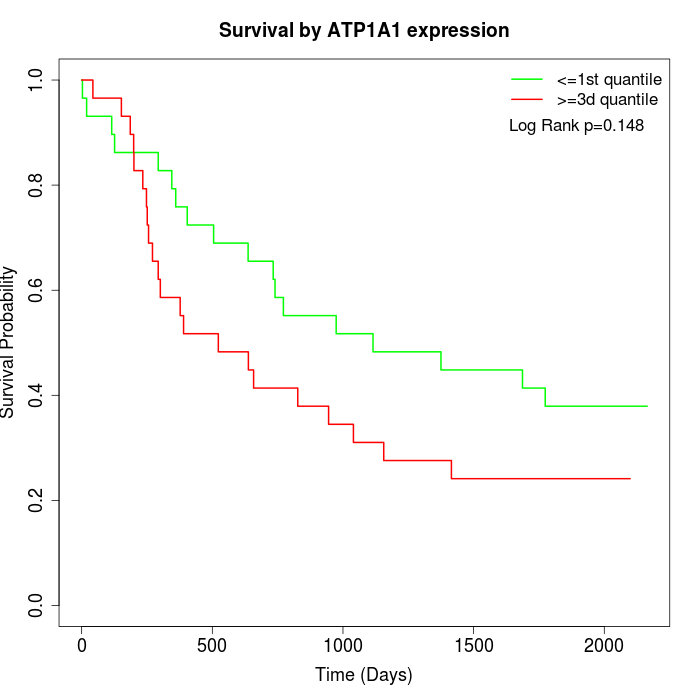
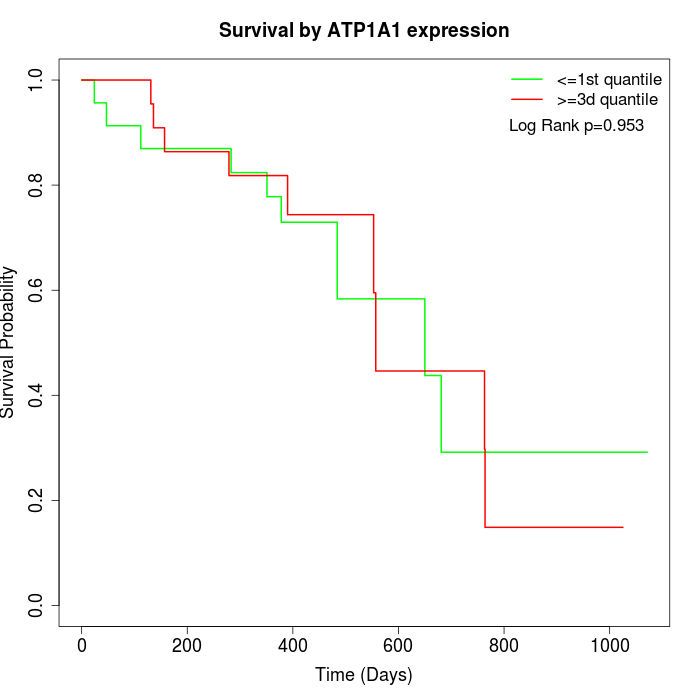
Survival by ATP1A1 expression:
Note: Click image to view full size file.
Copy number change of ATP1A1:
| Dataset | Gene | EntrezID | Gain | Loss | Normal | Detail |
|---|---|---|---|---|---|---|
| GSE15526 | ATP1A1 | 476 | 0 | 9 | 21 | |
| GSE20123 | ATP1A1 | 476 | 0 | 9 | 21 | |
| GSE43470 | ATP1A1 | 476 | 0 | 6 | 37 | |
| GSE46452 | ATP1A1 | 476 | 2 | 1 | 56 | |
| GSE47630 | ATP1A1 | 476 | 9 | 5 | 26 | |
| GSE54993 | ATP1A1 | 476 | 0 | 1 | 69 | |
| GSE54994 | ATP1A1 | 476 | 7 | 3 | 43 | |
| GSE60625 | ATP1A1 | 476 | 0 | 0 | 11 | |
| GSE74703 | ATP1A1 | 476 | 0 | 5 | 31 | |
| GSE74704 | ATP1A1 | 476 | 0 | 5 | 15 | |
| TCGA | ATP1A1 | 476 | 15 | 28 | 53 |
Total number of gains: 33; Total number of losses: 72; Total Number of normals: 383.
Somatic mutations of ATP1A1:
Generating mutation plots.
Highly correlated genes for ATP1A1:
Showing top 20/815 corelated genes with mean PCC>0.5.
| Gene1 | Gene2 | Mean PCC | Num. Datasets | Num. PCC<0 | Num. PCC>0.5 |
|---|---|---|---|---|---|
| ATP1A1 | CDC27 | 0.748185 | 3 | 0 | 3 |
| ATP1A1 | PRELID1 | 0.742609 | 3 | 0 | 3 |
| ATP1A1 | FAM89A | 0.719012 | 3 | 0 | 3 |
| ATP1A1 | EBNA1BP2 | 0.712813 | 3 | 0 | 3 |
| ATP1A1 | SEPHS1 | 0.7065 | 3 | 0 | 3 |
| ATP1A1 | RPS4X | 0.706399 | 3 | 0 | 3 |
| ATP1A1 | UCK2 | 0.69893 | 3 | 0 | 3 |
| ATP1A1 | AFAP1L2 | 0.6932 | 3 | 0 | 3 |
| ATP1A1 | HIBADH | 0.691772 | 3 | 0 | 3 |
| ATP1A1 | HM13 | 0.688666 | 3 | 0 | 3 |
| ATP1A1 | FYTTD1 | 0.681477 | 3 | 0 | 3 |
| ATP1A1 | NIT1 | 0.67851 | 3 | 0 | 3 |
| ATP1A1 | NOP56 | 0.678019 | 3 | 0 | 3 |
| ATP1A1 | PYCR2 | 0.673842 | 3 | 0 | 3 |
| ATP1A1 | ADCY3 | 0.673247 | 3 | 0 | 3 |
| ATP1A1 | NLN | 0.671659 | 3 | 0 | 3 |
| ATP1A1 | MET | 0.670125 | 6 | 0 | 5 |
| ATP1A1 | RMI2 | 0.668202 | 4 | 0 | 4 |
| ATP1A1 | DCAF13 | 0.667843 | 5 | 0 | 5 |
| ATP1A1 | PPAT | 0.66599 | 3 | 0 | 3 |
For details and further investigation, click here