| Full name: B melanoma antigen | Alias Symbol: CT2.1|BAGE1 | ||
| Type: protein-coding gene | Cytoband: 21p11.1 not on reference assembly | ||
| Entrez ID: 574 | HGNC ID: HGNC:942 | Ensembl Gene: | OMIM ID: 605167 |
Expression of BAGE:
| Dataset | Gene | EntrezID | Probe | Log2FC | Adj.pValue | Expression |
|---|---|---|---|---|---|---|
| GSE17351 | BAGE | 574 | 1555603_at | -0.0108 | 0.9687 | |
| GSE20347 | BAGE | 574 | 207712_at | 0.0297 | 0.5621 | |
| GSE23400 | BAGE | 574 | 207712_at | 0.0019 | 0.9251 | |
| GSE26886 | BAGE | 574 | 1555605_x_at | 0.0426 | 0.7168 | |
| GSE29001 | BAGE | 574 | 207712_at | 0.0433 | 0.7062 | |
| GSE38129 | BAGE | 574 | 207712_at | 0.0248 | 0.6532 | |
| GSE45670 | BAGE | 574 | 207712_at | 0.0426 | 0.6052 | |
| GSE63941 | BAGE | 574 | 207712_at | 0.6503 | 0.6719 | |
| GSE77861 | BAGE | 574 | 1555605_x_at | -0.0715 | 0.5375 | |
| GSE97050 | BAGE | 574 | A_33_P3366859 | -0.2211 | 0.3363 | |
| SRP133303 | BAGE | 574 | RNAseq | 1.0065 | 0.0014 | |
| TCGA | BAGE | 574 | RNAseq | 3.2837 | 0.0521 |
Upregulated datasets: 1; Downregulated datasets: 0.
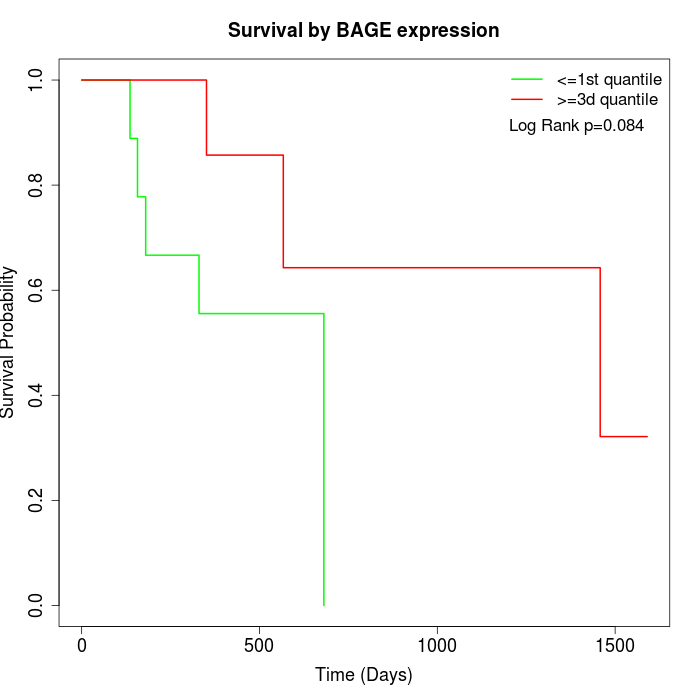
Survival by BAGE expression:
Note: Click image to view full size file.
Copy number change of BAGE:
| Dataset | Gene | EntrezID | Gain | Loss | Normal | Detail |
|---|---|---|---|---|---|---|
| GSE15526 | BAGE | 574 | 2 | 9 | 19 | |
| GSE20123 | BAGE | 574 | 2 | 9 | 19 | |
| GSE43470 | BAGE | 574 | 1 | 13 | 29 | |
| GSE46452 | BAGE | 574 | 1 | 22 | 36 | |
| GSE47630 | BAGE | 574 | 5 | 21 | 14 | |
| GSE54993 | BAGE | 574 | 10 | 1 | 59 | |
| GSE54994 | BAGE | 574 | 3 | 10 | 40 | |
| GSE60625 | BAGE | 574 | 0 | 0 | 11 | |
| GSE74703 | BAGE | 574 | 1 | 11 | 24 | |
| GSE74704 | BAGE | 574 | 2 | 6 | 12 |
Total number of gains: 27; Total number of losses: 102; Total Number of normals: 263.
Somatic mutations of BAGE:
Generating mutation plots.
Highly correlated genes for BAGE:
Showing top 20/106 corelated genes with mean PCC>0.5.
| Gene1 | Gene2 | Mean PCC | Num. Datasets | Num. PCC<0 | Num. PCC>0.5 |
|---|---|---|---|---|---|
| BAGE | LIPF | 0.77619 | 3 | 0 | 3 |
| BAGE | HK1 | 0.769285 | 3 | 0 | 3 |
| BAGE | NPFFR1 | 0.767377 | 3 | 0 | 3 |
| BAGE | NELL1 | 0.736846 | 3 | 0 | 3 |
| BAGE | NFAM1 | 0.712668 | 3 | 0 | 3 |
| BAGE | ZNF467 | 0.710869 | 4 | 0 | 3 |
| BAGE | PPP1R37 | 0.70216 | 4 | 0 | 3 |
| BAGE | KRT71 | 0.700534 | 3 | 0 | 3 |
| BAGE | CLC | 0.699413 | 4 | 0 | 4 |
| BAGE | AMBRA1 | 0.681929 | 4 | 0 | 4 |
| BAGE | KCNJ5 | 0.680984 | 3 | 0 | 3 |
| BAGE | TNFRSF4 | 0.678734 | 4 | 0 | 4 |
| BAGE | ATP1A4 | 0.678405 | 3 | 0 | 3 |
| BAGE | PSKH1 | 0.674374 | 3 | 0 | 3 |
| BAGE | ATP6AP1 | 0.672488 | 3 | 0 | 3 |
| BAGE | CCDC144A | 0.671837 | 4 | 0 | 3 |
| BAGE | CELA3A | 0.670373 | 4 | 0 | 3 |
| BAGE | IL34 | 0.66309 | 3 | 0 | 3 |
| BAGE | SHANK3 | 0.661891 | 3 | 0 | 3 |
| BAGE | FGFR1 | 0.660728 | 3 | 0 | 3 |
For details and further investigation, click here