| Full name: NFAT activating protein with ITAM motif 1 | Alias Symbol: CNAIP | ||
| Type: protein-coding gene | Cytoband: 22q13.2 | ||
| Entrez ID: 150372 | HGNC ID: HGNC:29872 | Ensembl Gene: ENSG00000235568 | OMIM ID: 608740 |
Screen Evidence:
| |||
Expression of NFAM1:
| Dataset | Gene | EntrezID | Probe | Log2FC | Adj.pValue | Expression |
|---|---|---|---|---|---|---|
| GSE17351 | NFAM1 | 150372 | 230322_at | 0.1966 | 0.5201 | |
| GSE26886 | NFAM1 | 150372 | 230322_at | 0.2087 | 0.0809 | |
| GSE45670 | NFAM1 | 150372 | 230322_at | 0.1645 | 0.0658 | |
| GSE63941 | NFAM1 | 150372 | 230322_at | 0.2889 | 0.0655 | |
| GSE77861 | NFAM1 | 150372 | 230322_at | 0.0618 | 0.7303 | |
| GSE97050 | NFAM1 | 150372 | A_33_P3292769 | -0.0453 | 0.8608 | |
| SRP007169 | NFAM1 | 150372 | RNAseq | 0.6838 | 0.4564 | |
| SRP064894 | NFAM1 | 150372 | RNAseq | 1.0002 | 0.0080 | |
| SRP133303 | NFAM1 | 150372 | RNAseq | 0.2256 | 0.3012 | |
| SRP159526 | NFAM1 | 150372 | RNAseq | -0.0701 | 0.9042 | |
| SRP193095 | NFAM1 | 150372 | RNAseq | 0.8421 | 0.0000 | |
| SRP219564 | NFAM1 | 150372 | RNAseq | 1.8042 | 0.0015 | |
| TCGA | NFAM1 | 150372 | RNAseq | 0.1082 | 0.5009 |
Upregulated datasets: 2; Downregulated datasets: 0.
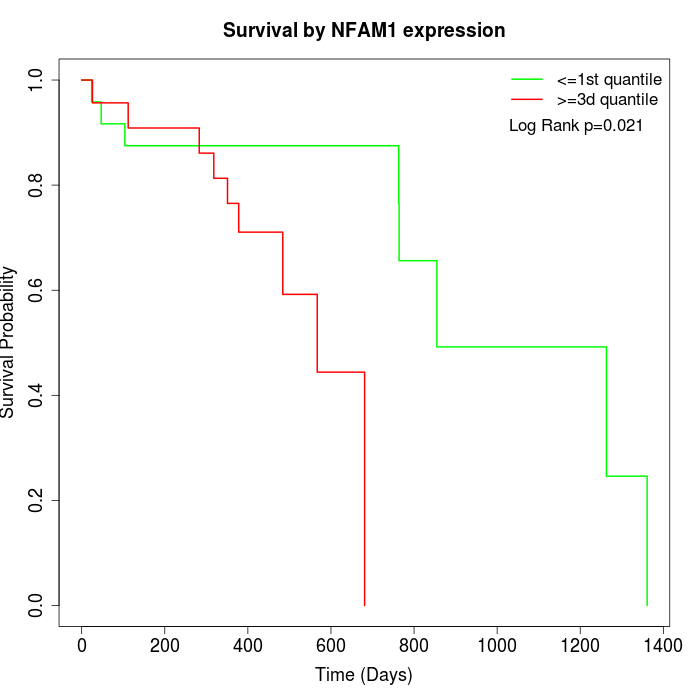
Survival by NFAM1 expression:
Note: Click image to view full size file.
Copy number change of NFAM1:
| Dataset | Gene | EntrezID | Gain | Loss | Normal | Detail |
|---|---|---|---|---|---|---|
| GSE15526 | NFAM1 | 150372 | 6 | 6 | 18 | |
| GSE20123 | NFAM1 | 150372 | 6 | 5 | 19 | |
| GSE43470 | NFAM1 | 150372 | 5 | 6 | 32 | |
| GSE46452 | NFAM1 | 150372 | 31 | 2 | 26 | |
| GSE47630 | NFAM1 | 150372 | 9 | 4 | 27 | |
| GSE54993 | NFAM1 | 150372 | 3 | 6 | 61 | |
| GSE54994 | NFAM1 | 150372 | 11 | 8 | 34 | |
| GSE60625 | NFAM1 | 150372 | 5 | 0 | 6 | |
| GSE74703 | NFAM1 | 150372 | 5 | 4 | 27 | |
| GSE74704 | NFAM1 | 150372 | 3 | 3 | 14 | |
| TCGA | NFAM1 | 150372 | 28 | 13 | 55 |
Total number of gains: 112; Total number of losses: 57; Total Number of normals: 319.
Somatic mutations of NFAM1:
Generating mutation plots.
Highly correlated genes for NFAM1:
Showing top 20/336 corelated genes with mean PCC>0.5.
| Gene1 | Gene2 | Mean PCC | Num. Datasets | Num. PCC<0 | Num. PCC>0.5 |
|---|---|---|---|---|---|
| NFAM1 | IL3RA | 0.819588 | 3 | 0 | 3 |
| NFAM1 | MAST1 | 0.803012 | 3 | 0 | 3 |
| NFAM1 | LIN37 | 0.78236 | 3 | 0 | 3 |
| NFAM1 | EXTL1 | 0.781529 | 3 | 0 | 3 |
| NFAM1 | BCAS4 | 0.767339 | 4 | 0 | 4 |
| NFAM1 | FN3K | 0.762557 | 3 | 0 | 3 |
| NFAM1 | RPTOR | 0.753216 | 3 | 0 | 3 |
| NFAM1 | BCORL1 | 0.751254 | 3 | 0 | 3 |
| NFAM1 | ADAMTS10 | 0.743111 | 3 | 0 | 3 |
| NFAM1 | KCNC4 | 0.739792 | 3 | 0 | 3 |
| NFAM1 | CASKIN2 | 0.739649 | 3 | 0 | 3 |
| NFAM1 | LGI2 | 0.737691 | 3 | 0 | 3 |
| NFAM1 | MIF | 0.734502 | 3 | 0 | 3 |
| NFAM1 | FNDC8 | 0.733916 | 3 | 0 | 3 |
| NFAM1 | MRPL38 | 0.731589 | 3 | 0 | 3 |
| NFAM1 | ZNF579 | 0.731586 | 3 | 0 | 3 |
| NFAM1 | MAPRE3 | 0.730582 | 4 | 0 | 4 |
| NFAM1 | SLC11A1 | 0.727858 | 3 | 0 | 3 |
| NFAM1 | KLF14 | 0.726205 | 3 | 0 | 3 |
| NFAM1 | KRT81 | 0.725753 | 3 | 0 | 3 |
For details and further investigation, click here