| Full name: BMP/retinoic acid inducible neural specific 3 | Alias Symbol: DBCCR1L|DBCCR1L1 | ||
| Type: protein-coding gene | Cytoband: 1q31.1 | ||
| Entrez ID: 339479 | HGNC ID: HGNC:22393 | Ensembl Gene: ENSG00000162670 | OMIM ID: 618390 |
Expression of BRINP3:
| Dataset | Gene | EntrezID | Probe | Log2FC | Adj.pValue | Expression |
|---|---|---|---|---|---|---|
| GSE17351 | BRINP3 | 339479 | 217562_at | -0.3612 | 0.8162 | |
| GSE20347 | BRINP3 | 339479 | 217562_at | -0.1370 | 0.3717 | |
| GSE23400 | BRINP3 | 339479 | 217562_at | -0.0721 | 0.0629 | |
| GSE26886 | BRINP3 | 339479 | 217562_at | 0.7255 | 0.1757 | |
| GSE29001 | BRINP3 | 339479 | 217562_at | 0.2194 | 0.6470 | |
| GSE38129 | BRINP3 | 339479 | 217562_at | -0.4328 | 0.0873 | |
| GSE45670 | BRINP3 | 339479 | 217562_at | -0.4966 | 0.1827 | |
| GSE53622 | BRINP3 | 339479 | 15795 | 0.1799 | 0.2583 | |
| GSE53624 | BRINP3 | 339479 | 15795 | 0.4003 | 0.0027 | |
| GSE63941 | BRINP3 | 339479 | 217562_at | -0.2620 | 0.1284 | |
| GSE77861 | BRINP3 | 339479 | 217562_at | -0.1458 | 0.1185 |
Upregulated datasets: 0; Downregulated datasets: 0.
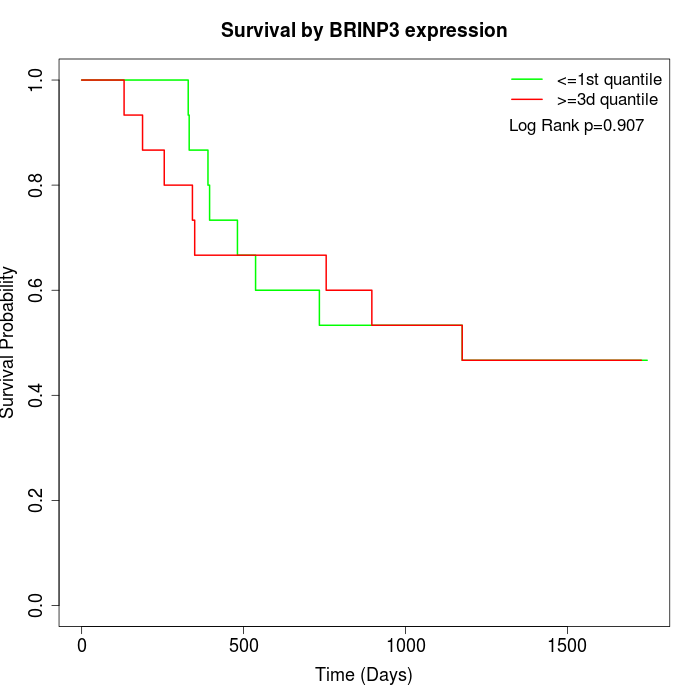
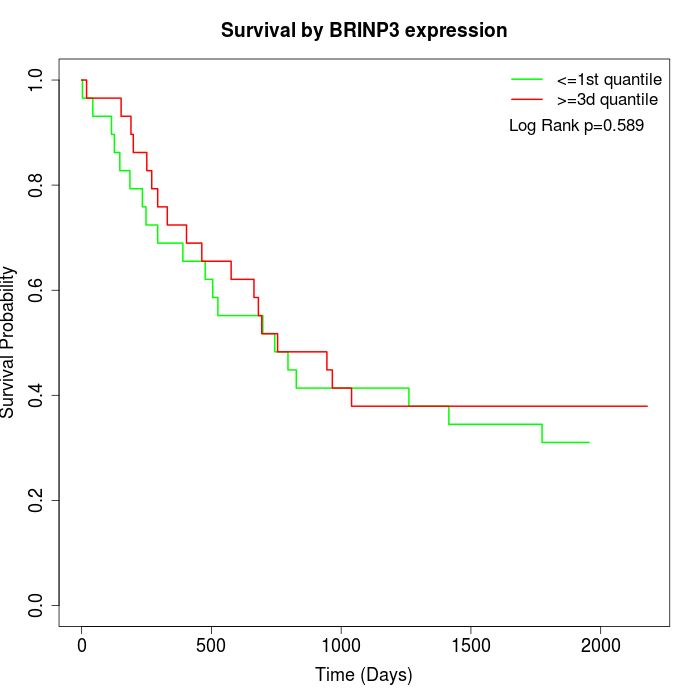
Survival by BRINP3 expression:
Note: Click image to view full size file.
Copy number change of BRINP3:
| Dataset | Gene | EntrezID | Gain | Loss | Normal | Detail |
|---|---|---|---|---|---|---|
| GSE15526 | BRINP3 | 339479 | 7 | 2 | 21 | |
| GSE20123 | BRINP3 | 339479 | 7 | 2 | 21 | |
| GSE43470 | BRINP3 | 339479 | 6 | 1 | 36 | |
| GSE46452 | BRINP3 | 339479 | 3 | 1 | 55 | |
| GSE47630 | BRINP3 | 339479 | 14 | 0 | 26 | |
| GSE54993 | BRINP3 | 339479 | 0 | 6 | 64 | |
| GSE54994 | BRINP3 | 339479 | 15 | 0 | 38 | |
| GSE60625 | BRINP3 | 339479 | 0 | 0 | 11 | |
| GSE74703 | BRINP3 | 339479 | 6 | 1 | 29 | |
| GSE74704 | BRINP3 | 339479 | 2 | 2 | 16 |
Total number of gains: 60; Total number of losses: 15; Total Number of normals: 317.
Somatic mutations of BRINP3:
Generating mutation plots.
Highly correlated genes for BRINP3:
Showing top 20/256 corelated genes with mean PCC>0.5.
| Gene1 | Gene2 | Mean PCC | Num. Datasets | Num. PCC<0 | Num. PCC>0.5 |
|---|---|---|---|---|---|
| BRINP3 | ILDR2 | 0.775752 | 3 | 0 | 3 |
| BRINP3 | SH3GL2 | 0.736858 | 7 | 0 | 7 |
| BRINP3 | GPM6A | 0.734698 | 6 | 0 | 4 |
| BRINP3 | WFDC3 | 0.716356 | 4 | 0 | 3 |
| BRINP3 | VIP | 0.715237 | 6 | 0 | 6 |
| BRINP3 | BEX1 | 0.712741 | 4 | 0 | 3 |
| BRINP3 | PDZRN4 | 0.705512 | 6 | 0 | 5 |
| BRINP3 | ESRRG | 0.69643 | 5 | 0 | 5 |
| BRINP3 | CDHR3 | 0.695632 | 3 | 0 | 3 |
| BRINP3 | PCDH9 | 0.693285 | 7 | 0 | 5 |
| BRINP3 | NCAM1 | 0.692412 | 5 | 0 | 4 |
| BRINP3 | KCNMB1 | 0.688893 | 4 | 0 | 4 |
| BRINP3 | PLEKHS1 | 0.688707 | 3 | 0 | 3 |
| BRINP3 | NLGN4X | 0.688052 | 3 | 0 | 3 |
| BRINP3 | ADAMTSL3 | 0.68796 | 5 | 0 | 5 |
| BRINP3 | GPAT2 | 0.683108 | 3 | 0 | 3 |
| BRINP3 | PYGM | 0.678154 | 5 | 0 | 4 |
| BRINP3 | HTR2B | 0.673879 | 6 | 0 | 4 |
| BRINP3 | EPHA3 | 0.673464 | 4 | 0 | 4 |
| BRINP3 | MYOM1 | 0.673185 | 6 | 0 | 5 |
For details and further investigation, click here