| Full name: chromatin target of PRMT1 | Alias Symbol: DKFZP547E1010|SRAG|FOP | ||
| Type: protein-coding gene | Cytoband: 1q21.3 | ||
| Entrez ID: 26097 | HGNC ID: HGNC:24511 | Ensembl Gene: ENSG00000160679 | OMIM ID: 614206 |
| Drug and gene relationship at DGIdb | |||
Screen Evidence:
| |||
Expression of CHTOP:
| Dataset | Gene | EntrezID | Probe | Log2FC | Adj.pValue | Expression |
|---|---|---|---|---|---|---|
| GSE17351 | CHTOP | 26097 | 202559_x_at | 0.3764 | 0.0943 | |
| GSE20347 | CHTOP | 26097 | 202559_x_at | 0.2286 | 0.0139 | |
| GSE23400 | CHTOP | 26097 | 202559_x_at | 0.1737 | 0.0001 | |
| GSE26886 | CHTOP | 26097 | 202559_x_at | -0.2667 | 0.0437 | |
| GSE29001 | CHTOP | 26097 | 202560_s_at | 0.1951 | 0.4229 | |
| GSE38129 | CHTOP | 26097 | 202559_x_at | 0.2184 | 0.0056 | |
| GSE45670 | CHTOP | 26097 | 202559_x_at | 0.2688 | 0.0011 | |
| GSE53622 | CHTOP | 26097 | 4301 | 0.0554 | 0.2557 | |
| GSE53624 | CHTOP | 26097 | 4301 | 0.1477 | 0.0019 | |
| GSE63941 | CHTOP | 26097 | 202559_x_at | 1.1509 | 0.0001 | |
| GSE77861 | CHTOP | 26097 | 202559_x_at | 0.2461 | 0.0461 | |
| GSE97050 | CHTOP | 26097 | A_33_P3243957 | 0.1161 | 0.7214 | |
| SRP007169 | CHTOP | 26097 | RNAseq | -0.3206 | 0.3419 | |
| SRP008496 | CHTOP | 26097 | RNAseq | -0.1534 | 0.3935 | |
| SRP064894 | CHTOP | 26097 | RNAseq | 0.1429 | 0.3589 | |
| SRP133303 | CHTOP | 26097 | RNAseq | 0.0591 | 0.7006 | |
| SRP159526 | CHTOP | 26097 | RNAseq | 0.2395 | 0.2098 | |
| SRP193095 | CHTOP | 26097 | RNAseq | -0.0588 | 0.4011 | |
| SRP219564 | CHTOP | 26097 | RNAseq | 0.3272 | 0.3187 |
Upregulated datasets: 1; Downregulated datasets: 0.
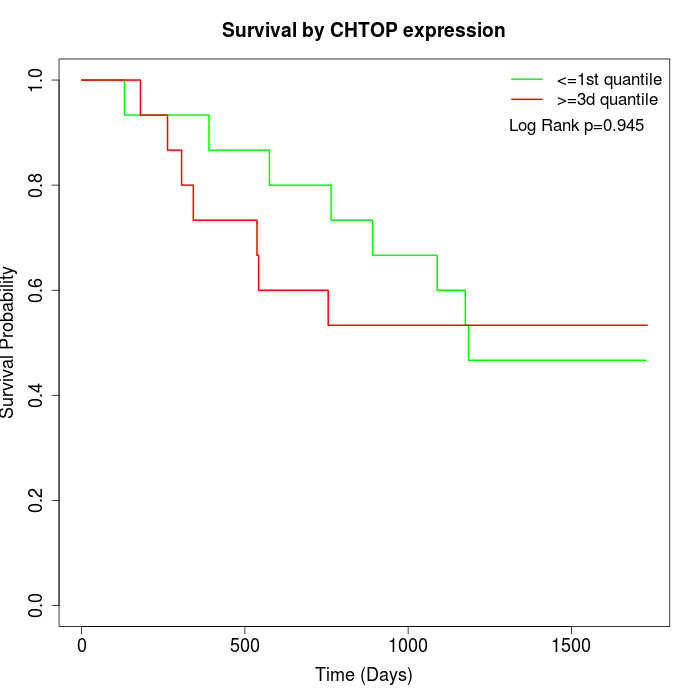
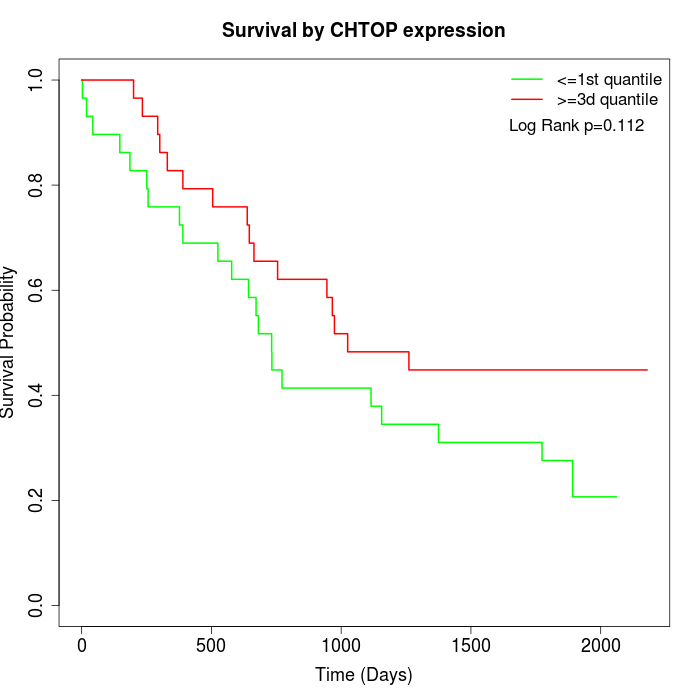
Survival by CHTOP expression:
Note: Click image to view full size file.
Copy number change of CHTOP:
| Dataset | Gene | EntrezID | Gain | Loss | Normal | Detail |
|---|---|---|---|---|---|---|
| GSE15526 | CHTOP | 26097 | 14 | 0 | 16 | |
| GSE20123 | CHTOP | 26097 | 14 | 0 | 16 | |
| GSE43470 | CHTOP | 26097 | 7 | 2 | 34 | |
| GSE46452 | CHTOP | 26097 | 2 | 1 | 56 | |
| GSE47630 | CHTOP | 26097 | 14 | 0 | 26 | |
| GSE54993 | CHTOP | 26097 | 0 | 4 | 66 | |
| GSE54994 | CHTOP | 26097 | 16 | 0 | 37 | |
| GSE60625 | CHTOP | 26097 | 0 | 0 | 11 | |
| GSE74703 | CHTOP | 26097 | 6 | 1 | 29 | |
| GSE74704 | CHTOP | 26097 | 7 | 0 | 13 | |
| TCGA | CHTOP | 26097 | 38 | 2 | 56 |
Total number of gains: 118; Total number of losses: 10; Total Number of normals: 360.
Somatic mutations of CHTOP:
Generating mutation plots.
Highly correlated genes for CHTOP:
Showing top 20/537 corelated genes with mean PCC>0.5.
| Gene1 | Gene2 | Mean PCC | Num. Datasets | Num. PCC<0 | Num. PCC>0.5 |
|---|---|---|---|---|---|
| CHTOP | PPP1CA | 0.756946 | 3 | 0 | 3 |
| CHTOP | SAPCD2 | 0.737184 | 3 | 0 | 3 |
| CHTOP | PLEKHH1 | 0.70046 | 3 | 0 | 3 |
| CHTOP | ZNF518B | 0.688668 | 4 | 0 | 3 |
| CHTOP | STX16 | 0.681561 | 3 | 0 | 3 |
| CHTOP | B3GNT5 | 0.681186 | 3 | 0 | 3 |
| CHTOP | MTBP | 0.674433 | 4 | 0 | 4 |
| CHTOP | CCDC18 | 0.673213 | 3 | 0 | 3 |
| CHTOP | CCDC85C | 0.67291 | 4 | 0 | 4 |
| CHTOP | FBXO11 | 0.67046 | 3 | 0 | 3 |
| CHTOP | USP37 | 0.665519 | 3 | 0 | 3 |
| CHTOP | CNOT10 | 0.665304 | 3 | 0 | 3 |
| CHTOP | PIP4K2C | 0.664849 | 3 | 0 | 3 |
| CHTOP | SDHAF2 | 0.66452 | 3 | 0 | 3 |
| CHTOP | SAMD1 | 0.66342 | 4 | 0 | 3 |
| CHTOP | DHX35 | 0.662928 | 3 | 0 | 3 |
| CHTOP | AMMECR1L | 0.658986 | 4 | 0 | 4 |
| CHTOP | LITAF | 0.65661 | 3 | 0 | 3 |
| CHTOP | PPIE | 0.655091 | 3 | 0 | 3 |
| CHTOP | ATXN7L3 | 0.655077 | 3 | 0 | 3 |
For details and further investigation, click here