| Full name: cytochrome P450 family 27 subfamily B member 1 | Alias Symbol: CYP1|P450c1 | ||
| Type: protein-coding gene | Cytoband: 12q14.1 | ||
| Entrez ID: 1594 | HGNC ID: HGNC:2606 | Ensembl Gene: ENSG00000111012 | OMIM ID: 609506 |
| Related drugs: CHEMBL255088, DECITABINE, INTERFERON GAMA-1B, TRICHOSTATIN... [more] | |||
CYP27B1 involved pathways:
| KEGG pathway | Description | View |
|---|---|---|
| hsa05152 | Tuberculosis |
Expression of CYP27B1:
| Dataset | Gene | EntrezID | Probe | Log2FC | Adj.pValue | Expression |
|---|---|---|---|---|---|---|
| GSE17351 | CYP27B1 | 1594 | 205676_at | 1.8599 | 0.0167 | |
| GSE20347 | CYP27B1 | 1594 | 205676_at | 0.7027 | 0.0125 | |
| GSE23400 | CYP27B1 | 1594 | 205676_at | 0.4097 | 0.0000 | |
| GSE26886 | CYP27B1 | 1594 | 205676_at | 0.5025 | 0.1012 | |
| GSE29001 | CYP27B1 | 1594 | 205676_at | 0.7236 | 0.0014 | |
| GSE38129 | CYP27B1 | 1594 | 205676_at | 1.0728 | 0.0000 | |
| GSE45670 | CYP27B1 | 1594 | 205676_at | 1.0971 | 0.0063 | |
| GSE53622 | CYP27B1 | 1594 | 37033 | 2.2752 | 0.0000 | |
| GSE53624 | CYP27B1 | 1594 | 37033 | 2.0245 | 0.0000 | |
| GSE63941 | CYP27B1 | 1594 | 205676_at | 0.7599 | 0.0339 | |
| GSE77861 | CYP27B1 | 1594 | 205676_at | 0.2985 | 0.0315 | |
| GSE97050 | CYP27B1 | 1594 | A_23_P36397 | 0.1909 | 0.5101 | |
| SRP007169 | CYP27B1 | 1594 | RNAseq | 2.9314 | 0.0023 | |
| SRP133303 | CYP27B1 | 1594 | RNAseq | 2.2710 | 0.0000 | |
| SRP159526 | CYP27B1 | 1594 | RNAseq | 2.1136 | 0.0001 | |
| SRP193095 | CYP27B1 | 1594 | RNAseq | 2.7493 | 0.0000 | |
| SRP219564 | CYP27B1 | 1594 | RNAseq | 0.9483 | 0.1826 | |
| TCGA | CYP27B1 | 1594 | RNAseq | 1.1817 | 0.0000 |
Upregulated datasets: 10; Downregulated datasets: 0.
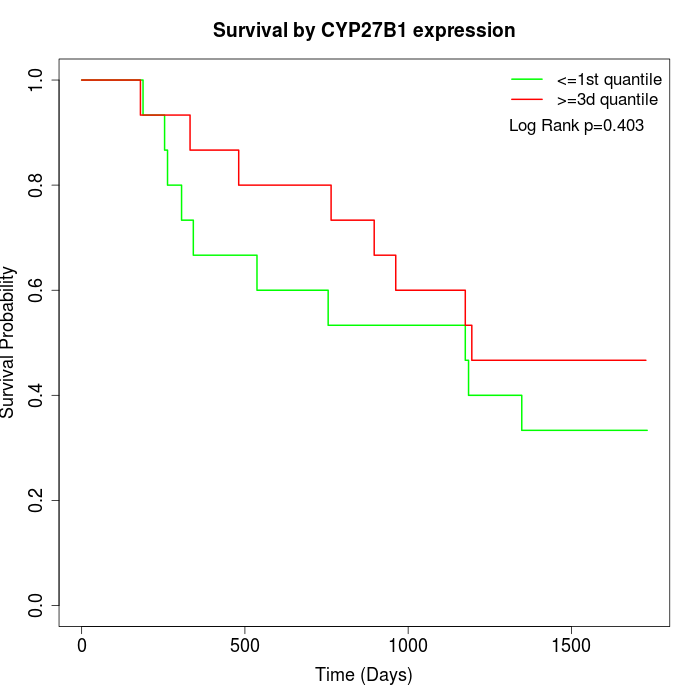
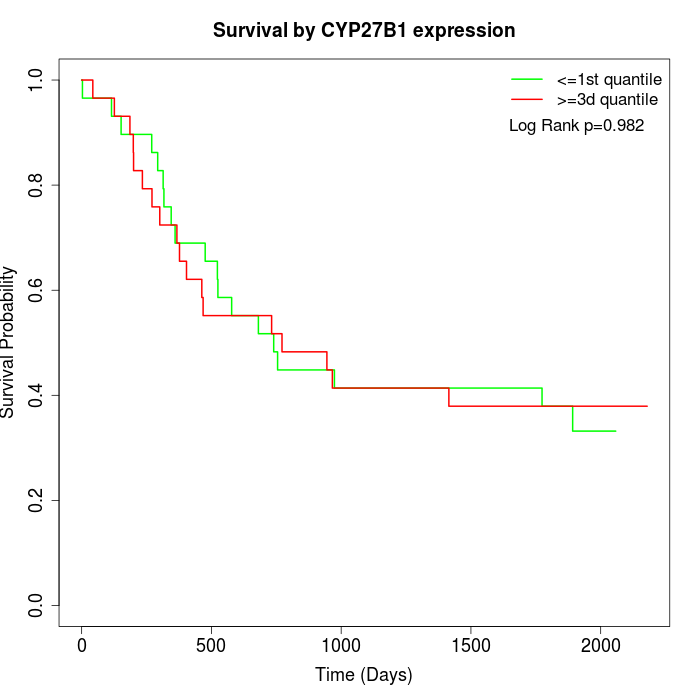
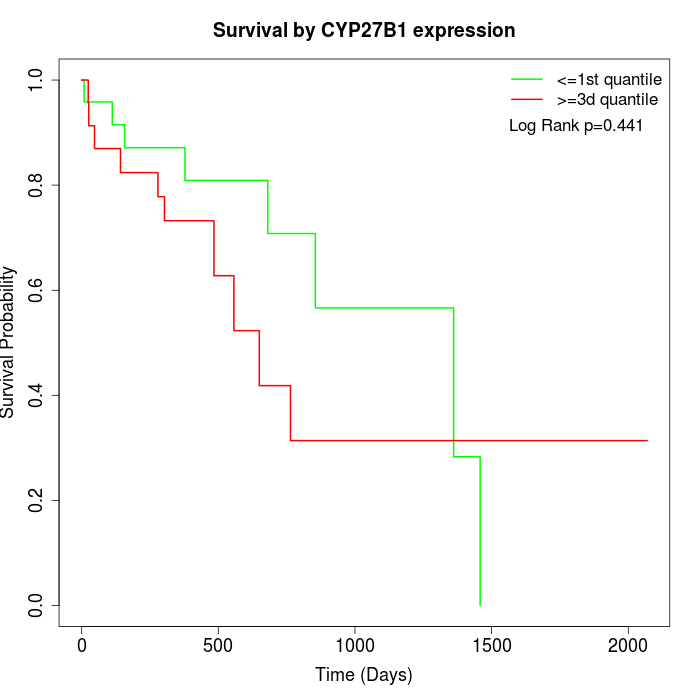
Survival by CYP27B1 expression:
Note: Click image to view full size file.
Copy number change of CYP27B1:
| Dataset | Gene | EntrezID | Gain | Loss | Normal | Detail |
|---|---|---|---|---|---|---|
| GSE15526 | CYP27B1 | 1594 | 7 | 1 | 22 | |
| GSE20123 | CYP27B1 | 1594 | 7 | 0 | 23 | |
| GSE43470 | CYP27B1 | 1594 | 2 | 0 | 41 | |
| GSE46452 | CYP27B1 | 1594 | 8 | 1 | 50 | |
| GSE47630 | CYP27B1 | 1594 | 10 | 2 | 28 | |
| GSE54993 | CYP27B1 | 1594 | 0 | 5 | 65 | |
| GSE54994 | CYP27B1 | 1594 | 4 | 1 | 48 | |
| GSE60625 | CYP27B1 | 1594 | 0 | 0 | 11 | |
| GSE74703 | CYP27B1 | 1594 | 2 | 0 | 34 | |
| GSE74704 | CYP27B1 | 1594 | 5 | 0 | 15 | |
| TCGA | CYP27B1 | 1594 | 14 | 10 | 72 |
Total number of gains: 59; Total number of losses: 20; Total Number of normals: 409.
Somatic mutations of CYP27B1:
Generating mutation plots.
Highly correlated genes for CYP27B1:
Showing top 20/958 corelated genes with mean PCC>0.5.
| Gene1 | Gene2 | Mean PCC | Num. Datasets | Num. PCC<0 | Num. PCC>0.5 |
|---|---|---|---|---|---|
| CYP27B1 | IL24 | 0.729415 | 9 | 0 | 9 |
| CYP27B1 | CMTM4 | 0.721545 | 3 | 0 | 3 |
| CYP27B1 | TMEM138 | 0.710124 | 5 | 0 | 5 |
| CYP27B1 | STRIP2 | 0.70232 | 6 | 0 | 6 |
| CYP27B1 | ADAM8 | 0.688524 | 9 | 0 | 9 |
| CYP27B1 | HOXC9 | 0.672757 | 4 | 0 | 3 |
| CYP27B1 | NTMT1 | 0.669848 | 5 | 0 | 4 |
| CYP27B1 | ITGA3 | 0.669351 | 12 | 0 | 10 |
| CYP27B1 | C1QTNF6 | 0.661432 | 5 | 0 | 4 |
| CYP27B1 | LAMC2 | 0.661406 | 12 | 0 | 11 |
| CYP27B1 | CDH3 | 0.659298 | 10 | 0 | 9 |
| CYP27B1 | DGKH | 0.658659 | 3 | 0 | 3 |
| CYP27B1 | ITGB4 | 0.65785 | 12 | 0 | 10 |
| CYP27B1 | GPNMB | 0.657839 | 4 | 0 | 3 |
| CYP27B1 | NUDCD1 | 0.65763 | 6 | 0 | 5 |
| CYP27B1 | SLC39A10 | 0.65613 | 4 | 0 | 3 |
| CYP27B1 | TRMT6 | 0.654939 | 6 | 0 | 5 |
| CYP27B1 | FLVCR1 | 0.654707 | 4 | 0 | 3 |
| CYP27B1 | PLA2G7 | 0.653502 | 11 | 0 | 11 |
| CYP27B1 | GBP5 | 0.653334 | 3 | 0 | 3 |
For details and further investigation, click here