| Full name: engulfment and cell motility 1 | Alias Symbol: KIAA0281|CED12|ELMO-1|CED-12 | ||
| Type: protein-coding gene | Cytoband: 7p14.2-p14.1 | ||
| Entrez ID: 9844 | HGNC ID: HGNC:16286 | Ensembl Gene: ENSG00000155849 | OMIM ID: 606420 |
| Drug and gene relationship at DGIdb | |||
ELMO1 involved pathways:
| KEGG pathway | Description | View |
|---|---|---|
| hsa04062 | Chemokine signaling pathway | |
| hsa05100 | Bacterial invasion of epithelial cells | |
| hsa05131 | Shigellosis |
Expression of ELMO1:
| Dataset | Gene | EntrezID | Probe | Log2FC | Adj.pValue | Expression |
|---|---|---|---|---|---|---|
| GSE17351 | ELMO1 | 9844 | 204513_s_at | -0.4521 | 0.6060 | |
| GSE20347 | ELMO1 | 9844 | 204513_s_at | 0.3504 | 0.0504 | |
| GSE23400 | ELMO1 | 9844 | 204513_s_at | 0.1681 | 0.0341 | |
| GSE26886 | ELMO1 | 9844 | 204513_s_at | -0.2534 | 0.2253 | |
| GSE29001 | ELMO1 | 9844 | 204513_s_at | 0.1727 | 0.5367 | |
| GSE38129 | ELMO1 | 9844 | 204513_s_at | 0.1836 | 0.5497 | |
| GSE45670 | ELMO1 | 9844 | 204513_s_at | -0.4179 | 0.1776 | |
| GSE53622 | ELMO1 | 9844 | 15891 | -0.3328 | 0.0151 | |
| GSE53624 | ELMO1 | 9844 | 15891 | 0.1488 | 0.2081 | |
| GSE63941 | ELMO1 | 9844 | 204513_s_at | 0.0377 | 0.9746 | |
| GSE77861 | ELMO1 | 9844 | 204513_s_at | 0.2913 | 0.2055 | |
| GSE97050 | ELMO1 | 9844 | A_23_P122937 | 0.3059 | 0.3221 | |
| SRP007169 | ELMO1 | 9844 | RNAseq | 0.6427 | 0.1227 | |
| SRP008496 | ELMO1 | 9844 | RNAseq | 0.8586 | 0.0081 | |
| SRP064894 | ELMO1 | 9844 | RNAseq | 0.5971 | 0.0512 | |
| SRP133303 | ELMO1 | 9844 | RNAseq | -0.0149 | 0.9517 | |
| SRP159526 | ELMO1 | 9844 | RNAseq | 1.7405 | 0.0038 | |
| SRP193095 | ELMO1 | 9844 | RNAseq | 0.8640 | 0.0000 | |
| SRP219564 | ELMO1 | 9844 | RNAseq | 0.5951 | 0.1217 | |
| TCGA | ELMO1 | 9844 | RNAseq | 0.0864 | 0.5029 |
Upregulated datasets: 1; Downregulated datasets: 0.
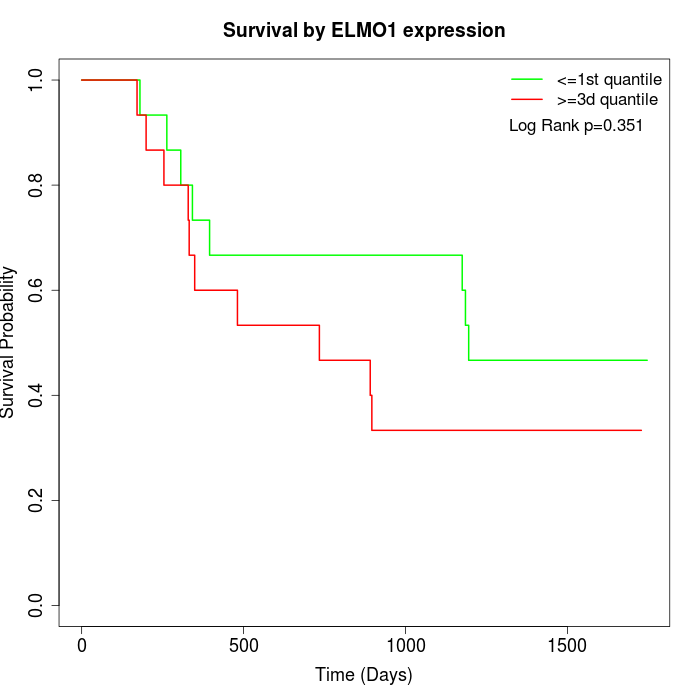
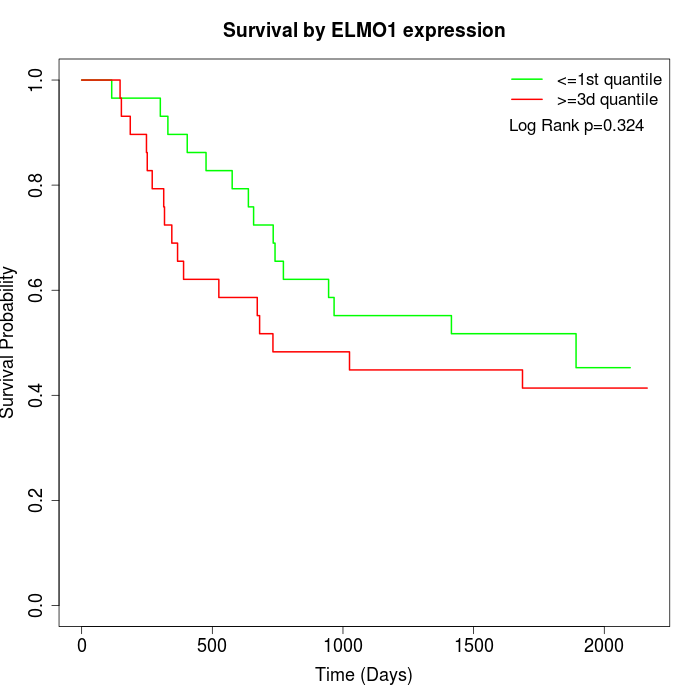
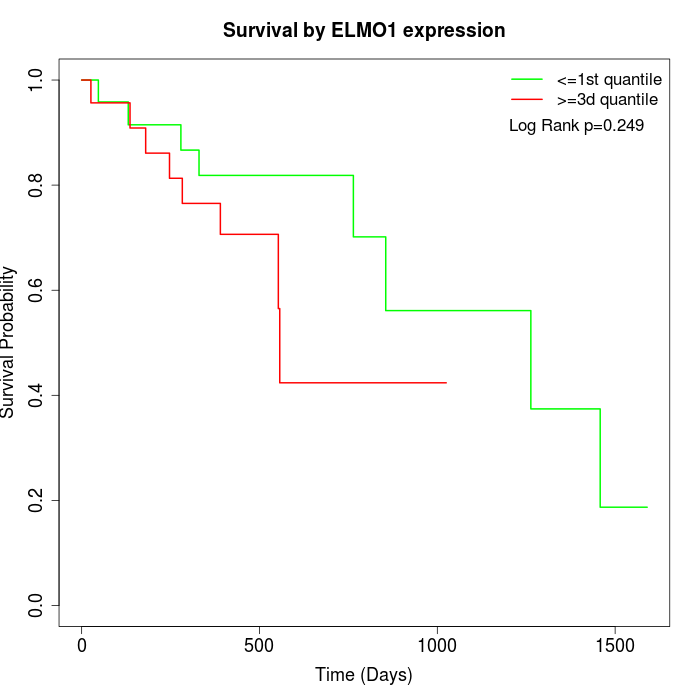
Survival by ELMO1 expression:
Note: Click image to view full size file.
Copy number change of ELMO1:
| Dataset | Gene | EntrezID | Gain | Loss | Normal | Detail |
|---|---|---|---|---|---|---|
| GSE15526 | ELMO1 | 9844 | 16 | 0 | 14 | |
| GSE20123 | ELMO1 | 9844 | 15 | 0 | 15 | |
| GSE43470 | ELMO1 | 9844 | 4 | 0 | 39 | |
| GSE46452 | ELMO1 | 9844 | 11 | 1 | 47 | |
| GSE47630 | ELMO1 | 9844 | 8 | 1 | 31 | |
| GSE54993 | ELMO1 | 9844 | 0 | 7 | 63 | |
| GSE54994 | ELMO1 | 9844 | 18 | 3 | 32 | |
| GSE60625 | ELMO1 | 9844 | 0 | 0 | 11 | |
| GSE74703 | ELMO1 | 9844 | 4 | 0 | 32 | |
| GSE74704 | ELMO1 | 9844 | 8 | 0 | 12 | |
| TCGA | ELMO1 | 9844 | 54 | 6 | 36 |
Total number of gains: 138; Total number of losses: 18; Total Number of normals: 332.
Somatic mutations of ELMO1:
Generating mutation plots.
Highly correlated genes for ELMO1:
Showing top 20/580 corelated genes with mean PCC>0.5.
| Gene1 | Gene2 | Mean PCC | Num. Datasets | Num. PCC<0 | Num. PCC>0.5 |
|---|---|---|---|---|---|
| ELMO1 | CD151 | 0.794885 | 3 | 0 | 3 |
| ELMO1 | BMPR2 | 0.783261 | 3 | 0 | 3 |
| ELMO1 | ZFAND3 | 0.77022 | 3 | 0 | 3 |
| ELMO1 | PBX2 | 0.73694 | 3 | 0 | 3 |
| ELMO1 | GSX1 | 0.736703 | 3 | 0 | 3 |
| ELMO1 | SOX5 | 0.736024 | 3 | 0 | 3 |
| ELMO1 | PODN | 0.712767 | 3 | 0 | 3 |
| ELMO1 | TBC1D16 | 0.712229 | 3 | 0 | 3 |
| ELMO1 | SP4 | 0.705233 | 4 | 0 | 3 |
| ELMO1 | GNG2 | 0.703629 | 4 | 0 | 4 |
| ELMO1 | U2AF1 | 0.70302 | 3 | 0 | 3 |
| ELMO1 | IFIT1 | 0.698225 | 3 | 0 | 3 |
| ELMO1 | IFI6 | 0.696756 | 3 | 0 | 3 |
| ELMO1 | CDKN1C | 0.694468 | 4 | 0 | 3 |
| ELMO1 | TNFSF12 | 0.693746 | 3 | 0 | 3 |
| ELMO1 | R3HDM2 | 0.691394 | 3 | 0 | 3 |
| ELMO1 | RWDD2A | 0.689622 | 4 | 0 | 3 |
| ELMO1 | PTPN1 | 0.688963 | 3 | 0 | 3 |
| ELMO1 | CNRIP1 | 0.685059 | 6 | 0 | 5 |
| ELMO1 | SSH1 | 0.684322 | 4 | 0 | 3 |
For details and further investigation, click here