| Full name: long intergenic non-protein coding RNA 589 | Alias Symbol: TSLNC8 | ||
| Type: non-coding RNA | Cytoband: 8p12 | ||
| Entrez ID: 619351 | HGNC ID: HGNC:32299 | Ensembl Gene: ENSG00000251191 | OMIM ID: |
Expression of LINC00589:
| Dataset | Gene | EntrezID | Probe | Log2FC | Adj.pValue | Expression |
|---|---|---|---|---|---|---|
| GSE17351 | LINC00589 | 619351 | 232718_at | 0.0548 | 0.8752 | |
| GSE26886 | LINC00589 | 619351 | 232718_at | -0.0776 | 0.5738 | |
| GSE45670 | LINC00589 | 619351 | 232718_at | -0.0050 | 0.9647 | |
| GSE53622 | LINC00589 | 619351 | 36558 | 0.1258 | 0.1074 | |
| GSE53624 | LINC00589 | 619351 | 961 | 0.9184 | 0.0000 | |
| GSE63941 | LINC00589 | 619351 | 232718_at | 0.2753 | 0.2324 | |
| GSE77861 | LINC00589 | 619351 | 232718_at | -0.1951 | 0.0499 |
Upregulated datasets: 0; Downregulated datasets: 0.
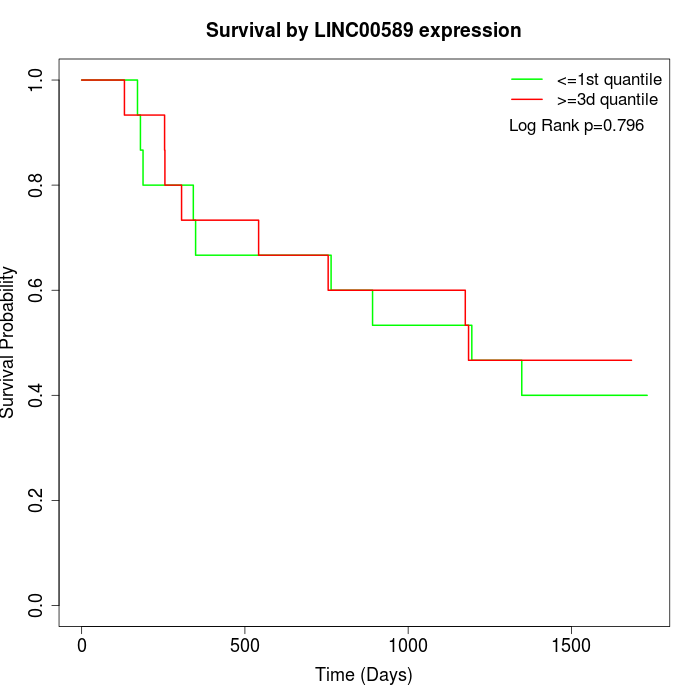
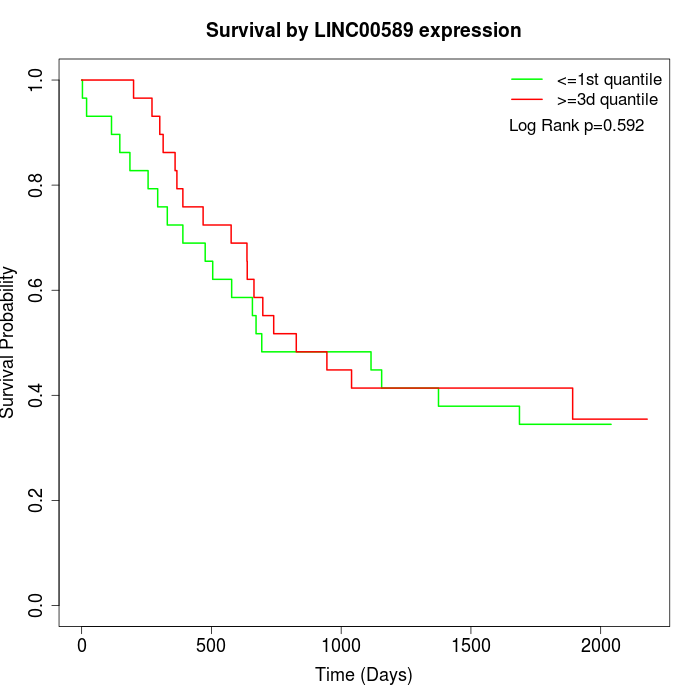
Survival by LINC00589 expression:
Note: Click image to view full size file.
Copy number change of LINC00589:
| Dataset | Gene | EntrezID | Gain | Loss | Normal | Detail |
|---|---|---|---|---|---|---|
| GSE15526 | LINC00589 | 619351 | 4 | 10 | 16 | |
| GSE20123 | LINC00589 | 619351 | 4 | 10 | 16 | |
| GSE43470 | LINC00589 | 619351 | 4 | 7 | 32 | |
| GSE46452 | LINC00589 | 619351 | 13 | 13 | 33 | |
| GSE47630 | LINC00589 | 619351 | 10 | 8 | 22 | |
| GSE54993 | LINC00589 | 619351 | 2 | 15 | 53 | |
| GSE54994 | LINC00589 | 619351 | 10 | 15 | 28 | |
| GSE60625 | LINC00589 | 619351 | 3 | 0 | 8 | |
| GSE74703 | LINC00589 | 619351 | 4 | 6 | 26 | |
| GSE74704 | LINC00589 | 619351 | 3 | 7 | 10 | |
| TCGA | LINC00589 | 619351 | 18 | 40 | 38 |
Total number of gains: 75; Total number of losses: 131; Total Number of normals: 282.
Somatic mutations of LINC00589:
Generating mutation plots.
Highly correlated genes for LINC00589:
Showing top 20/34 corelated genes with mean PCC>0.5.
| Gene1 | Gene2 | Mean PCC | Num. Datasets | Num. PCC<0 | Num. PCC>0.5 |
|---|---|---|---|---|---|
| LINC00589 | KIF12 | 0.618875 | 3 | 0 | 3 |
| LINC00589 | RD3 | 0.610681 | 3 | 0 | 3 |
| LINC00589 | KCNQ1 | 0.602569 | 3 | 0 | 3 |
| LINC00589 | ERVV-1 | 0.592691 | 4 | 0 | 3 |
| LINC00589 | SLC6A7 | 0.589841 | 3 | 0 | 3 |
| LINC00589 | DEFB118 | 0.586558 | 3 | 0 | 3 |
| LINC00589 | FANCD2OS | 0.583008 | 4 | 0 | 3 |
| LINC00589 | HCN1 | 0.582747 | 4 | 0 | 3 |
| LINC00589 | FATE1 | 0.576888 | 5 | 0 | 3 |
| LINC00589 | KLK4 | 0.569731 | 4 | 0 | 3 |
| LINC00589 | DEFB119 | 0.569528 | 4 | 0 | 3 |
| LINC00589 | SPEF2 | 0.569513 | 3 | 0 | 3 |
| LINC00589 | PIP | 0.568781 | 3 | 0 | 3 |
| LINC00589 | ENPP3 | 0.568074 | 4 | 0 | 3 |
| LINC00589 | MS4A15 | 0.566166 | 3 | 0 | 3 |
| LINC00589 | LILRA2 | 0.563485 | 4 | 0 | 3 |
| LINC00589 | NRAP | 0.559312 | 4 | 0 | 3 |
| LINC00589 | LINC01210 | 0.554642 | 3 | 0 | 3 |
| LINC00589 | PRSS55 | 0.550869 | 4 | 0 | 3 |
| LINC00589 | E2F4 | 0.550831 | 4 | 0 | 3 |
For details and further investigation, click here