| Full name: long intergenic non-protein coding RNA 702 | Alias Symbol: | ||
| Type: non-coding RNA | Cytoband: 10p15.1 | ||
| Entrez ID: 100652988 | HGNC ID: HGNC:44676 | Ensembl Gene: ENSG00000233117 | OMIM ID: |
Expression of LINC00702:
| Dataset | Gene | EntrezID | Probe | Log2FC | Adj.pValue | Expression |
|---|---|---|---|---|---|---|
| GSE17351 | LINC00702 | 100652988 | 236304_at | -1.4078 | 0.4993 | |
| GSE26886 | LINC00702 | 100652988 | 236304_at | 0.6539 | 0.0353 | |
| GSE45670 | LINC00702 | 100652988 | 236304_at | -1.6735 | 0.0004 | |
| GSE53622 | LINC00702 | 100652988 | 69212 | -0.6401 | 0.0130 | |
| GSE53624 | LINC00702 | 100652988 | 69212 | 0.0086 | 0.9682 | |
| GSE63941 | LINC00702 | 100652988 | 236304_at | 0.0192 | 0.9012 | |
| GSE77861 | LINC00702 | 100652988 | 236304_at | 0.0353 | 0.6572 | |
| SRP133303 | LINC00702 | 100652988 | RNAseq | -0.1974 | 0.7926 |
Upregulated datasets: 0; Downregulated datasets: 1.
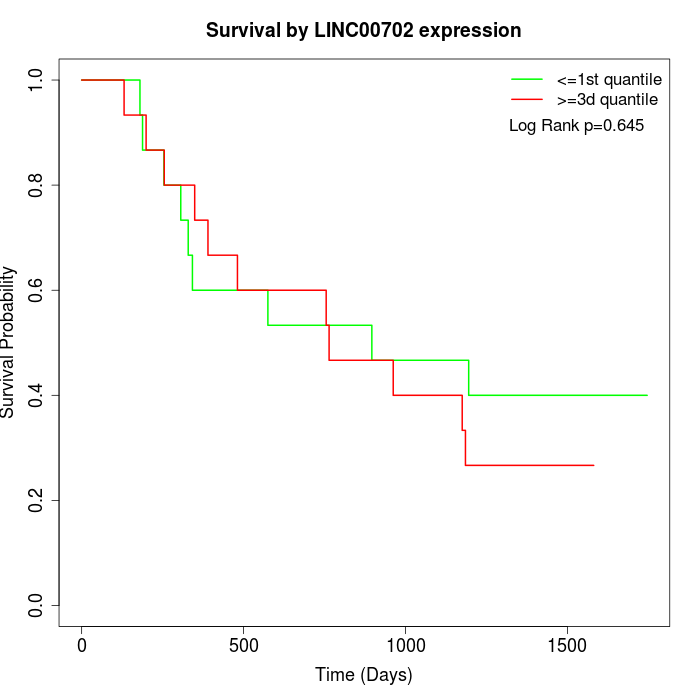
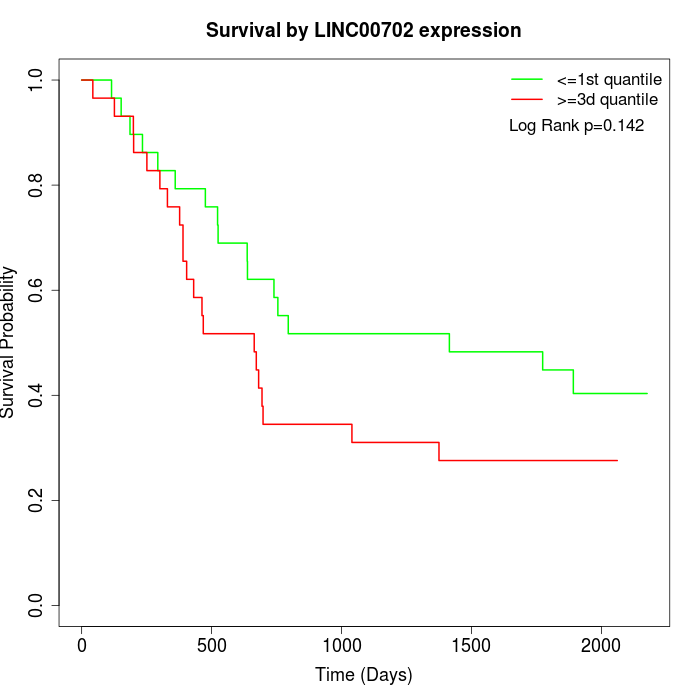
Survival by LINC00702 expression:
Note: Click image to view full size file.
Copy number change of LINC00702:
| Dataset | Gene | EntrezID | Gain | Loss | Normal | Detail |
|---|---|---|---|---|---|---|
| GSE15526 | LINC00702 | 100652988 | 5 | 8 | 17 | |
| GSE20123 | LINC00702 | 100652988 | 5 | 7 | 18 | |
| GSE43470 | LINC00702 | 100652988 | 3 | 6 | 34 | |
| GSE46452 | LINC00702 | 100652988 | 2 | 14 | 43 | |
| GSE47630 | LINC00702 | 100652988 | 6 | 14 | 20 | |
| GSE54993 | LINC00702 | 100652988 | 11 | 0 | 59 | |
| GSE54994 | LINC00702 | 100652988 | 4 | 10 | 39 | |
| GSE60625 | LINC00702 | 100652988 | 0 | 0 | 11 | |
| GSE74703 | LINC00702 | 100652988 | 2 | 4 | 30 | |
| GSE74704 | LINC00702 | 100652988 | 1 | 6 | 13 | |
| TCGA | LINC00702 | 100652988 | 19 | 25 | 52 |
Total number of gains: 58; Total number of losses: 94; Total Number of normals: 336.
Somatic mutations of LINC00702:
Generating mutation plots.
Highly correlated genes for LINC00702:
Showing top 20/346 corelated genes with mean PCC>0.5.
| Gene1 | Gene2 | Mean PCC | Num. Datasets | Num. PCC<0 | Num. PCC>0.5 |
|---|---|---|---|---|---|
| LINC00702 | MYOM1 | 0.782769 | 3 | 0 | 3 |
| LINC00702 | ITGBL1 | 0.772286 | 3 | 0 | 3 |
| LINC00702 | DENND2A | 0.76695 | 4 | 0 | 4 |
| LINC00702 | MITF | 0.763216 | 3 | 0 | 3 |
| LINC00702 | RBM24 | 0.751995 | 3 | 0 | 3 |
| LINC00702 | PYGM | 0.748195 | 3 | 0 | 3 |
| LINC00702 | CNGA3 | 0.738871 | 3 | 0 | 3 |
| LINC00702 | MYOC | 0.737779 | 3 | 0 | 3 |
| LINC00702 | CACNA1C | 0.737723 | 4 | 0 | 4 |
| LINC00702 | PPP3CB | 0.734954 | 3 | 0 | 3 |
| LINC00702 | ILK | 0.733922 | 4 | 0 | 4 |
| LINC00702 | HTR2B | 0.732901 | 3 | 0 | 3 |
| LINC00702 | PLP1 | 0.730161 | 3 | 0 | 3 |
| LINC00702 | KCNMB1 | 0.727267 | 3 | 0 | 3 |
| LINC00702 | HECTD2 | 0.725607 | 3 | 0 | 3 |
| LINC00702 | ITGA1 | 0.722396 | 4 | 0 | 4 |
| LINC00702 | RHOB | 0.72198 | 3 | 0 | 3 |
| LINC00702 | LHCGR | 0.72132 | 3 | 0 | 3 |
| LINC00702 | LRCH2 | 0.715507 | 4 | 0 | 4 |
| LINC00702 | PBXIP1 | 0.710593 | 3 | 0 | 3 |
For details and further investigation, click here