| Full name: lysophosphatidic acid receptor 6 | Alias Symbol: P2Y5 | ||
| Type: protein-coding gene | Cytoband: 13q14.2 | ||
| Entrez ID: 10161 | HGNC ID: HGNC:15520 | Ensembl Gene: ENSG00000139679 | OMIM ID: 609239 |
LPAR6 involved pathways:
| KEGG pathway | Description | View |
|---|---|---|
| hsa04072 | Phospholipase D signaling pathway | |
| hsa04151 | PI3K-Akt signaling pathway |
Expression of LPAR6:
| Dataset | Gene | EntrezID | Probe | Log2FC | Adj.pValue | Expression |
|---|---|---|---|---|---|---|
| GSE17351 | LPAR6 | 10161 | 218589_at | -1.1260 | 0.3753 | |
| GSE20347 | LPAR6 | 10161 | 218589_at | -1.4447 | 0.0005 | |
| GSE23400 | LPAR6 | 10161 | 218589_at | -0.6123 | 0.0001 | |
| GSE26886 | LPAR6 | 10161 | 218589_at | -1.8322 | 0.0001 | |
| GSE29001 | LPAR6 | 10161 | 218589_at | -1.3990 | 0.0021 | |
| GSE38129 | LPAR6 | 10161 | 218589_at | -0.8347 | 0.0310 | |
| GSE45670 | LPAR6 | 10161 | 218589_at | -0.3554 | 0.1727 | |
| GSE53622 | LPAR6 | 10161 | 25963 | -0.6704 | 0.0000 | |
| GSE53624 | LPAR6 | 10161 | 25963 | -1.2531 | 0.0000 | |
| GSE63941 | LPAR6 | 10161 | 218589_at | -1.5337 | 0.1713 | |
| GSE77861 | LPAR6 | 10161 | 218589_at | -0.7850 | 0.0017 | |
| GSE97050 | LPAR6 | 10161 | A_23_P2705 | 0.3141 | 0.5114 | |
| SRP007169 | LPAR6 | 10161 | RNAseq | -2.6920 | 0.0000 | |
| SRP008496 | LPAR6 | 10161 | RNAseq | -2.7060 | 0.0000 | |
| SRP064894 | LPAR6 | 10161 | RNAseq | -1.0116 | 0.0000 | |
| SRP133303 | LPAR6 | 10161 | RNAseq | -0.7656 | 0.0060 | |
| SRP159526 | LPAR6 | 10161 | RNAseq | -2.1332 | 0.0000 | |
| SRP193095 | LPAR6 | 10161 | RNAseq | -1.2736 | 0.0000 | |
| SRP219564 | LPAR6 | 10161 | RNAseq | -1.1238 | 0.0903 | |
| TCGA | LPAR6 | 10161 | RNAseq | 0.3663 | 0.0013 |
Upregulated datasets: 0; Downregulated datasets: 9.
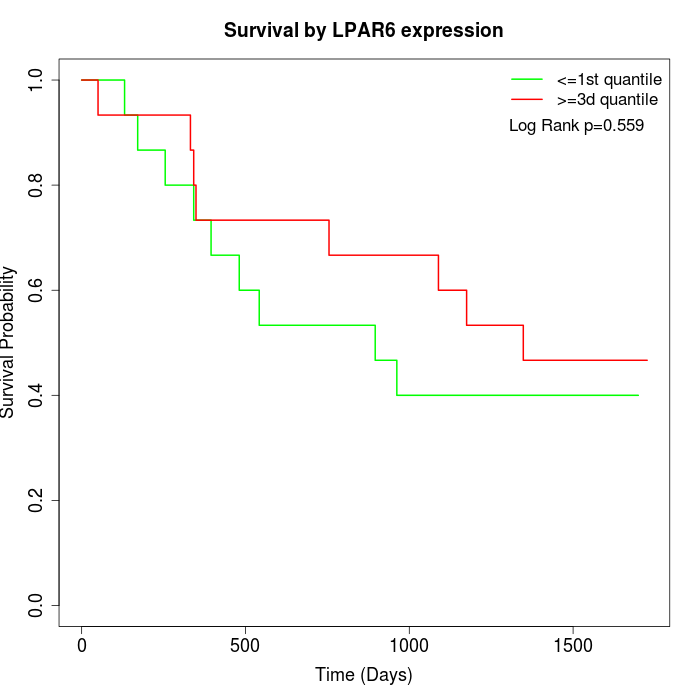
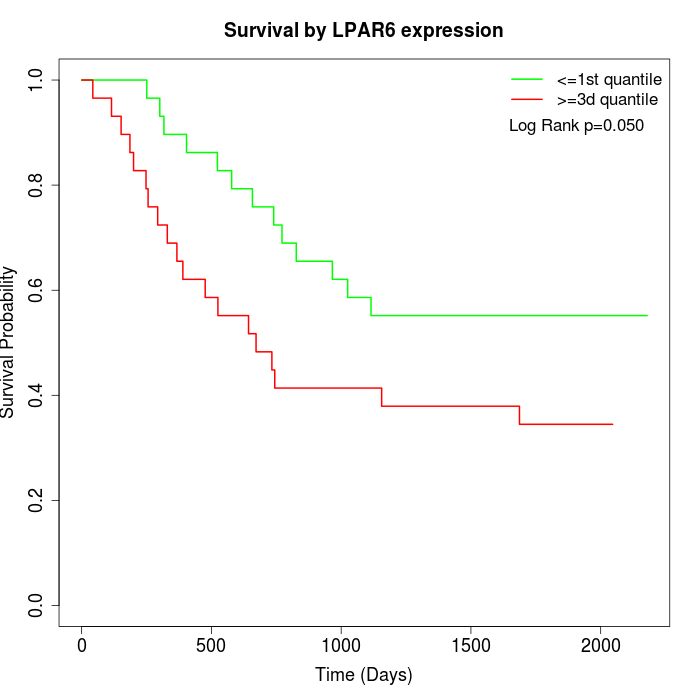
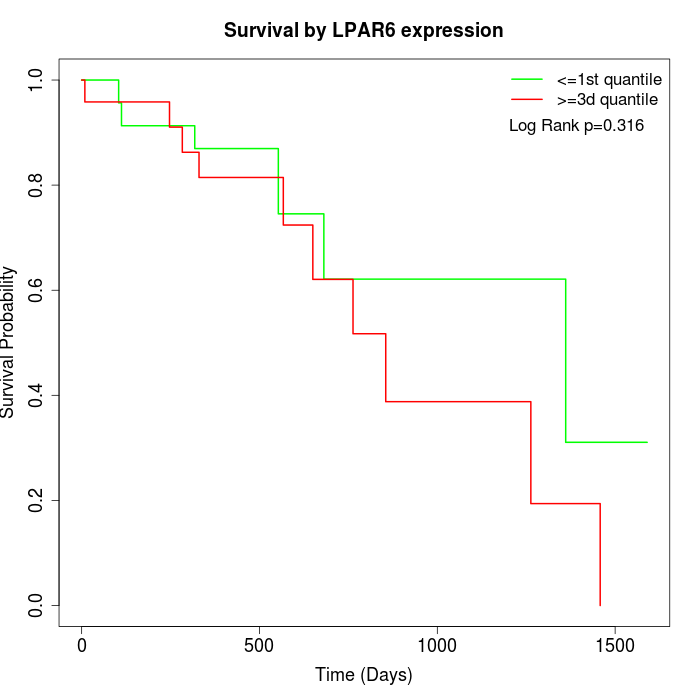
Survival by LPAR6 expression:
Note: Click image to view full size file.
Copy number change of LPAR6:
| Dataset | Gene | EntrezID | Gain | Loss | Normal | Detail |
|---|---|---|---|---|---|---|
| GSE15526 | LPAR6 | 10161 | 0 | 13 | 17 | |
| GSE20123 | LPAR6 | 10161 | 0 | 12 | 18 | |
| GSE43470 | LPAR6 | 10161 | 3 | 13 | 27 | |
| GSE46452 | LPAR6 | 10161 | 0 | 32 | 27 | |
| GSE47630 | LPAR6 | 10161 | 2 | 27 | 11 | |
| GSE54993 | LPAR6 | 10161 | 12 | 2 | 56 | |
| GSE54994 | LPAR6 | 10161 | 1 | 17 | 35 | |
| GSE60625 | LPAR6 | 10161 | 0 | 3 | 8 | |
| GSE74703 | LPAR6 | 10161 | 3 | 10 | 23 | |
| GSE74704 | LPAR6 | 10161 | 0 | 9 | 11 | |
| TCGA | LPAR6 | 10161 | 7 | 40 | 49 |
Total number of gains: 28; Total number of losses: 178; Total Number of normals: 282.
Somatic mutations of LPAR6:
Generating mutation plots.
Highly correlated genes for LPAR6:
Showing top 20/1097 corelated genes with mean PCC>0.5.
| Gene1 | Gene2 | Mean PCC | Num. Datasets | Num. PCC<0 | Num. PCC>0.5 |
|---|---|---|---|---|---|
| LPAR6 | FYTTD1 | 0.844227 | 3 | 0 | 3 |
| LPAR6 | DLL1 | 0.78709 | 3 | 0 | 3 |
| LPAR6 | STT3B | 0.764766 | 4 | 0 | 4 |
| LPAR6 | IER3IP1 | 0.749221 | 3 | 0 | 3 |
| LPAR6 | TMEM38A | 0.748469 | 3 | 0 | 3 |
| LPAR6 | AK2 | 0.734259 | 3 | 0 | 3 |
| LPAR6 | MLKL | 0.728298 | 4 | 0 | 4 |
| LPAR6 | RNF165 | 0.719788 | 3 | 0 | 3 |
| LPAR6 | ALG14 | 0.71361 | 3 | 0 | 3 |
| LPAR6 | DOCK8 | 0.705644 | 4 | 0 | 4 |
| LPAR6 | PAIP2 | 0.697548 | 3 | 0 | 3 |
| LPAR6 | SCARF1 | 0.691903 | 3 | 0 | 3 |
| LPAR6 | SLC35A4 | 0.691582 | 4 | 0 | 3 |
| LPAR6 | INPP1 | 0.69155 | 9 | 0 | 9 |
| LPAR6 | SYTL1 | 0.690498 | 6 | 0 | 6 |
| LPAR6 | PPP1R3D | 0.690288 | 4 | 0 | 3 |
| LPAR6 | PLEKHA7 | 0.689883 | 6 | 0 | 6 |
| LPAR6 | NUAK2 | 0.687683 | 3 | 0 | 3 |
| LPAR6 | CCNG2 | 0.687593 | 10 | 0 | 10 |
| LPAR6 | SEPSECS | 0.686933 | 3 | 0 | 3 |
For details and further investigation, click here