| Full name: methyltransferase like 22 | Alias Symbol: FLJ12433|MGC2654 | ||
| Type: protein-coding gene | Cytoband: 16p13.2 | ||
| Entrez ID: 79091 | HGNC ID: HGNC:28368 | Ensembl Gene: ENSG00000067365 | OMIM ID: 615261 |
| Drug and gene relationship at DGIdb | |||
Expression of METTL22:
| Dataset | Gene | EntrezID | Probe | Log2FC | Adj.pValue | Expression |
|---|---|---|---|---|---|---|
| GSE17351 | METTL22 | 79091 | 218945_at | -0.0478 | 0.9264 | |
| GSE20347 | METTL22 | 79091 | 218945_at | 0.1610 | 0.2476 | |
| GSE23400 | METTL22 | 79091 | 218945_at | 0.1345 | 0.0081 | |
| GSE26886 | METTL22 | 79091 | 218945_at | 0.6012 | 0.0069 | |
| GSE29001 | METTL22 | 79091 | 218945_at | 0.4885 | 0.0127 | |
| GSE38129 | METTL22 | 79091 | 218945_at | 0.1389 | 0.2163 | |
| GSE45670 | METTL22 | 79091 | 218945_at | 0.1661 | 0.3762 | |
| GSE53622 | METTL22 | 79091 | 29910 | 0.2500 | 0.0000 | |
| GSE53624 | METTL22 | 79091 | 29910 | 0.3278 | 0.0000 | |
| GSE63941 | METTL22 | 79091 | 218945_at | -0.7688 | 0.0409 | |
| GSE77861 | METTL22 | 79091 | 218945_at | 0.1746 | 0.2531 | |
| GSE97050 | METTL22 | 79091 | A_23_P15285 | 0.0418 | 0.8506 | |
| SRP007169 | METTL22 | 79091 | RNAseq | -0.0124 | 0.9767 | |
| SRP008496 | METTL22 | 79091 | RNAseq | 0.3513 | 0.2322 | |
| SRP064894 | METTL22 | 79091 | RNAseq | 0.4869 | 0.0017 | |
| SRP133303 | METTL22 | 79091 | RNAseq | 0.2820 | 0.0171 | |
| SRP159526 | METTL22 | 79091 | RNAseq | 0.3661 | 0.2335 | |
| SRP193095 | METTL22 | 79091 | RNAseq | 0.5318 | 0.0000 | |
| SRP219564 | METTL22 | 79091 | RNAseq | 0.1196 | 0.7458 |
Upregulated datasets: 0; Downregulated datasets: 0.
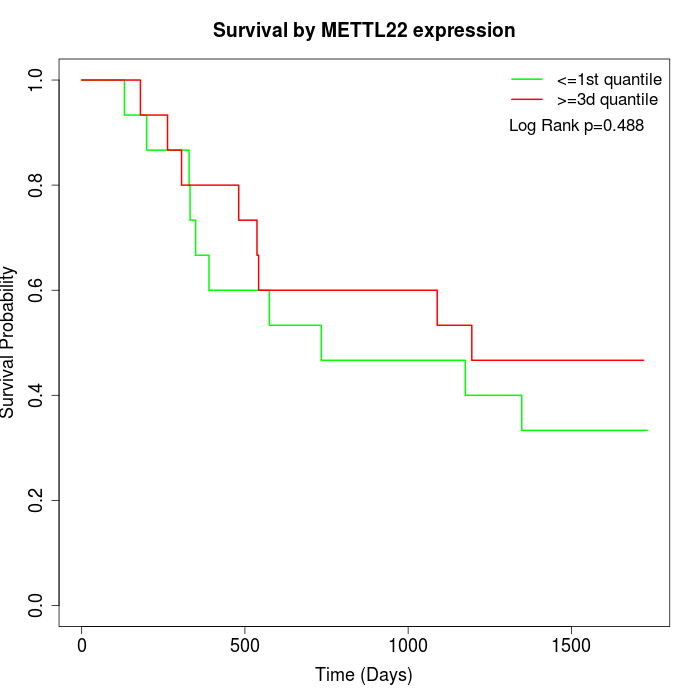
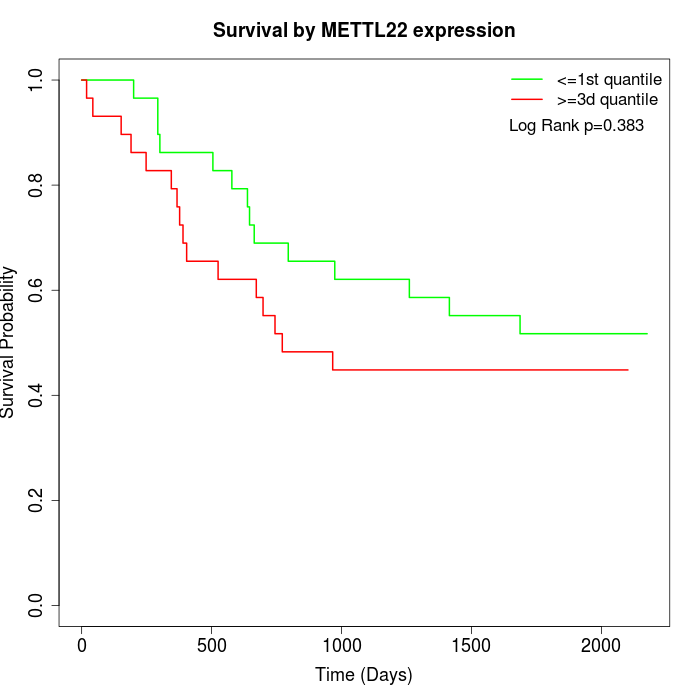
Survival by METTL22 expression:
Note: Click image to view full size file.
Copy number change of METTL22:
| Dataset | Gene | EntrezID | Gain | Loss | Normal | Detail |
|---|---|---|---|---|---|---|
| GSE15526 | METTL22 | 79091 | 4 | 4 | 22 | |
| GSE20123 | METTL22 | 79091 | 4 | 3 | 23 | |
| GSE43470 | METTL22 | 79091 | 3 | 2 | 38 | |
| GSE46452 | METTL22 | 79091 | 38 | 1 | 20 | |
| GSE47630 | METTL22 | 79091 | 13 | 6 | 21 | |
| GSE54993 | METTL22 | 79091 | 3 | 5 | 62 | |
| GSE54994 | METTL22 | 79091 | 5 | 9 | 39 | |
| GSE60625 | METTL22 | 79091 | 4 | 0 | 7 | |
| GSE74703 | METTL22 | 79091 | 3 | 2 | 31 | |
| GSE74704 | METTL22 | 79091 | 2 | 1 | 17 | |
| TCGA | METTL22 | 79091 | 20 | 13 | 63 |
Total number of gains: 99; Total number of losses: 46; Total Number of normals: 343.
Somatic mutations of METTL22:
Generating mutation plots.
Highly correlated genes for METTL22:
Showing top 20/128 corelated genes with mean PCC>0.5.
| Gene1 | Gene2 | Mean PCC | Num. Datasets | Num. PCC<0 | Num. PCC>0.5 |
|---|---|---|---|---|---|
| METTL22 | MRPL10 | 0.743056 | 3 | 0 | 3 |
| METTL22 | RNF25 | 0.710987 | 4 | 0 | 4 |
| METTL22 | BLOC1S6 | 0.693837 | 3 | 0 | 3 |
| METTL22 | NSMF | 0.644059 | 3 | 0 | 3 |
| METTL22 | OSTM1-AS1 | 0.610189 | 3 | 0 | 3 |
| METTL22 | C16orf72 | 0.605819 | 4 | 0 | 3 |
| METTL22 | BBS12 | 0.599495 | 3 | 0 | 3 |
| METTL22 | ADAM17 | 0.597134 | 4 | 0 | 3 |
| METTL22 | WDR24 | 0.594552 | 4 | 0 | 3 |
| METTL22 | SCAND1 | 0.593199 | 5 | 0 | 4 |
| METTL22 | PDE4A | 0.592387 | 3 | 0 | 3 |
| METTL22 | CCNA1 | 0.583217 | 6 | 0 | 4 |
| METTL22 | FBLIM1 | 0.582221 | 5 | 0 | 5 |
| METTL22 | KIF7 | 0.581278 | 4 | 0 | 3 |
| METTL22 | PHF20 | 0.5785 | 4 | 0 | 4 |
| METTL22 | CD2BP2 | 0.573593 | 5 | 0 | 4 |
| METTL22 | CREG2 | 0.570563 | 4 | 0 | 3 |
| METTL22 | ADSL | 0.570404 | 6 | 0 | 5 |
| METTL22 | ALDH1L2 | 0.569824 | 4 | 0 | 4 |
| METTL22 | RBM33 | 0.569191 | 4 | 0 | 3 |
For details and further investigation, click here