| Full name: NAD(P) dependent steroid dehydrogenase-like | Alias Symbol: XAP104|H105e3|SDR31E1 | ||
| Type: protein-coding gene | Cytoband: Xq28 | ||
| Entrez ID: 50814 | HGNC ID: HGNC:13398 | Ensembl Gene: ENSG00000147383 | OMIM ID: 300275 |
Expression of NSDHL:
| Dataset | Gene | EntrezID | Probe | Log2FC | Adj.pValue | Expression |
|---|---|---|---|---|---|---|
| GSE17351 | NSDHL | 50814 | 215093_at | -0.7008 | 0.2342 | |
| GSE20347 | NSDHL | 50814 | 215093_at | -0.5095 | 0.0015 | |
| GSE23400 | NSDHL | 50814 | 209279_s_at | -0.4255 | 0.0002 | |
| GSE26886 | NSDHL | 50814 | 215093_at | -0.1767 | 0.3120 | |
| GSE29001 | NSDHL | 50814 | 215093_at | -0.6114 | 0.0956 | |
| GSE38129 | NSDHL | 50814 | 215093_at | -0.1402 | 0.5588 | |
| GSE45670 | NSDHL | 50814 | 215093_at | 0.0860 | 0.6764 | |
| GSE53622 | NSDHL | 50814 | 115329 | -0.5160 | 0.0001 | |
| GSE53624 | NSDHL | 50814 | 115329 | -0.5169 | 0.0000 | |
| GSE63941 | NSDHL | 50814 | 215093_at | -0.4922 | 0.3687 | |
| GSE77861 | NSDHL | 50814 | 209279_s_at | -0.4348 | 0.0711 | |
| GSE97050 | NSDHL | 50814 | A_33_P3236082 | -0.0012 | 0.9977 | |
| SRP007169 | NSDHL | 50814 | RNAseq | -1.5955 | 0.0002 | |
| SRP008496 | NSDHL | 50814 | RNAseq | -1.7776 | 0.0000 | |
| SRP064894 | NSDHL | 50814 | RNAseq | -0.4908 | 0.0429 | |
| SRP133303 | NSDHL | 50814 | RNAseq | -0.0653 | 0.7998 | |
| SRP159526 | NSDHL | 50814 | RNAseq | -0.8760 | 0.0169 | |
| SRP193095 | NSDHL | 50814 | RNAseq | -0.9781 | 0.0000 | |
| SRP219564 | NSDHL | 50814 | RNAseq | -0.9132 | 0.0859 | |
| TCGA | NSDHL | 50814 | RNAseq | 0.2811 | 0.0001 |
Upregulated datasets: 0; Downregulated datasets: 2.
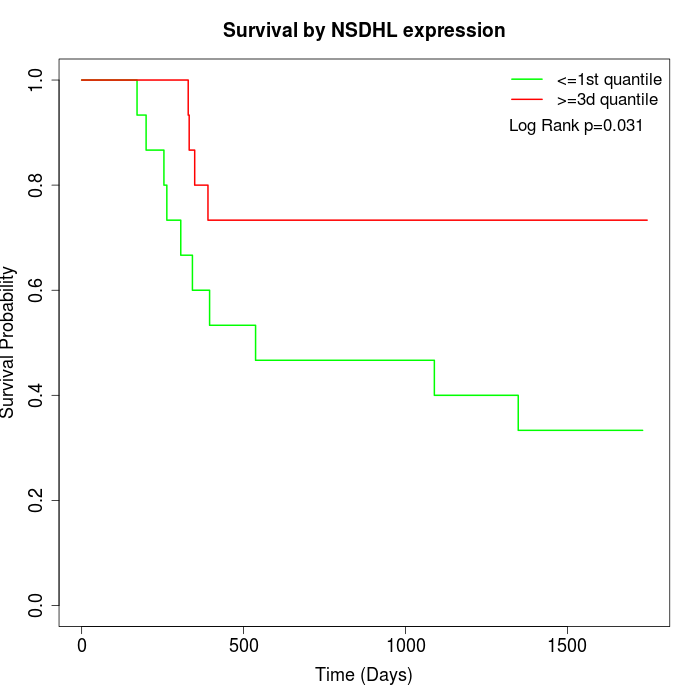
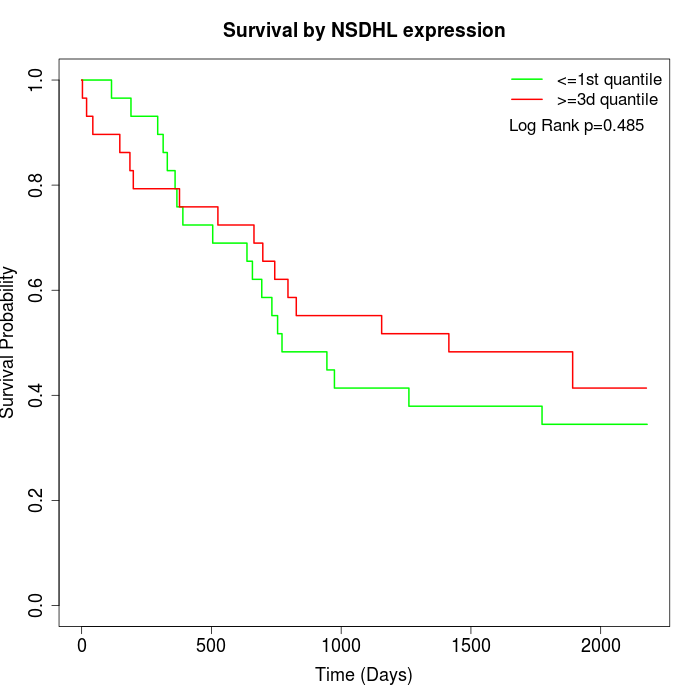
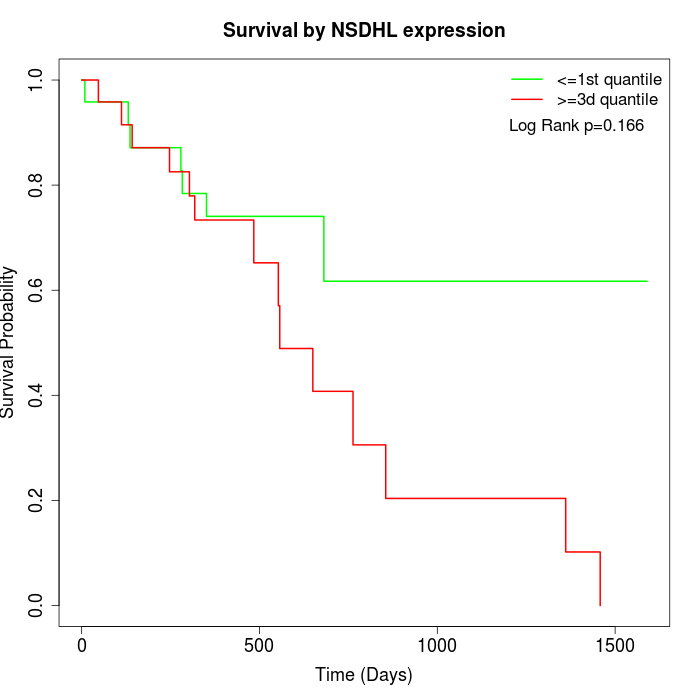
Survival by NSDHL expression:
Note: Click image to view full size file.
Copy number change of NSDHL:
No record found for this gene.
Somatic mutations of NSDHL:
Generating mutation plots.
Highly correlated genes for NSDHL:
Showing top 20/990 corelated genes with mean PCC>0.5.
| Gene1 | Gene2 | Mean PCC | Num. Datasets | Num. PCC<0 | Num. PCC>0.5 |
|---|---|---|---|---|---|
| NSDHL | NUAK2 | 0.784126 | 3 | 0 | 3 |
| NSDHL | CAPNS2 | 0.755937 | 4 | 0 | 4 |
| NSDHL | NIPAL1 | 0.753059 | 3 | 0 | 3 |
| NSDHL | S100A16 | 0.710997 | 7 | 0 | 6 |
| NSDHL | BLOC1S2 | 0.709067 | 3 | 0 | 3 |
| NSDHL | TMEM134 | 0.706457 | 8 | 0 | 7 |
| NSDHL | TALDO1 | 0.705851 | 5 | 0 | 5 |
| NSDHL | ARHGAP27 | 0.70506 | 6 | 0 | 5 |
| NSDHL | CERS3 | 0.69907 | 6 | 0 | 5 |
| NSDHL | EPGN | 0.698299 | 3 | 0 | 3 |
| NSDHL | KCNK6 | 0.696793 | 5 | 0 | 5 |
| NSDHL | EPHA1 | 0.692949 | 9 | 0 | 9 |
| NSDHL | PGM2 | 0.69182 | 7 | 0 | 6 |
| NSDHL | BTBD11 | 0.69106 | 7 | 0 | 6 |
| NSDHL | FBXL16 | 0.690297 | 6 | 0 | 6 |
| NSDHL | GNA15 | 0.688579 | 10 | 0 | 10 |
| NSDHL | LTA4H | 0.687858 | 3 | 0 | 3 |
| NSDHL | DUOXA1 | 0.687137 | 7 | 0 | 6 |
| NSDHL | TTC39B | 0.684714 | 3 | 0 | 3 |
| NSDHL | UPK3B | 0.684164 | 3 | 0 | 3 |
For details and further investigation, click here