| Full name: phospholipid scramblase 4 | Alias Symbol: | ||
| Type: protein-coding gene | Cytoband: 3q24 | ||
| Entrez ID: 57088 | HGNC ID: HGNC:16497 | Ensembl Gene: ENSG00000114698 | OMIM ID: 607612 |
Expression of PLSCR4:
| Dataset | Gene | EntrezID | Probe | Log2FC | Adj.pValue | Expression |
|---|---|---|---|---|---|---|
| GSE17351 | PLSCR4 | 57088 | 218901_at | -0.2686 | 0.6712 | |
| GSE20347 | PLSCR4 | 57088 | 218901_at | 0.4622 | 0.2164 | |
| GSE23400 | PLSCR4 | 57088 | 218901_at | 0.0534 | 0.7909 | |
| GSE26886 | PLSCR4 | 57088 | 218901_at | 0.1677 | 0.6884 | |
| GSE29001 | PLSCR4 | 57088 | 218901_at | 1.0675 | 0.0025 | |
| GSE38129 | PLSCR4 | 57088 | 218901_at | -0.2749 | 0.3926 | |
| GSE45670 | PLSCR4 | 57088 | 218901_at | -1.0308 | 0.0058 | |
| GSE53622 | PLSCR4 | 57088 | 441 | -0.6316 | 0.0001 | |
| GSE53624 | PLSCR4 | 57088 | 20481 | -0.4247 | 0.0001 | |
| GSE63941 | PLSCR4 | 57088 | 218901_at | -3.9216 | 0.0046 | |
| GSE77861 | PLSCR4 | 57088 | 218901_at | 0.3423 | 0.6426 | |
| GSE97050 | PLSCR4 | 57088 | A_23_P91910 | -0.5982 | 0.1453 | |
| SRP007169 | PLSCR4 | 57088 | RNAseq | 1.3208 | 0.0507 | |
| SRP008496 | PLSCR4 | 57088 | RNAseq | 1.3091 | 0.0053 | |
| SRP064894 | PLSCR4 | 57088 | RNAseq | -0.0687 | 0.8600 | |
| SRP133303 | PLSCR4 | 57088 | RNAseq | -0.6561 | 0.0198 | |
| SRP159526 | PLSCR4 | 57088 | RNAseq | -0.2801 | 0.4283 | |
| SRP193095 | PLSCR4 | 57088 | RNAseq | 0.7365 | 0.0018 | |
| SRP219564 | PLSCR4 | 57088 | RNAseq | -0.2953 | 0.5771 | |
| TCGA | PLSCR4 | 57088 | RNAseq | -0.1591 | 0.1751 |
Upregulated datasets: 2; Downregulated datasets: 2.
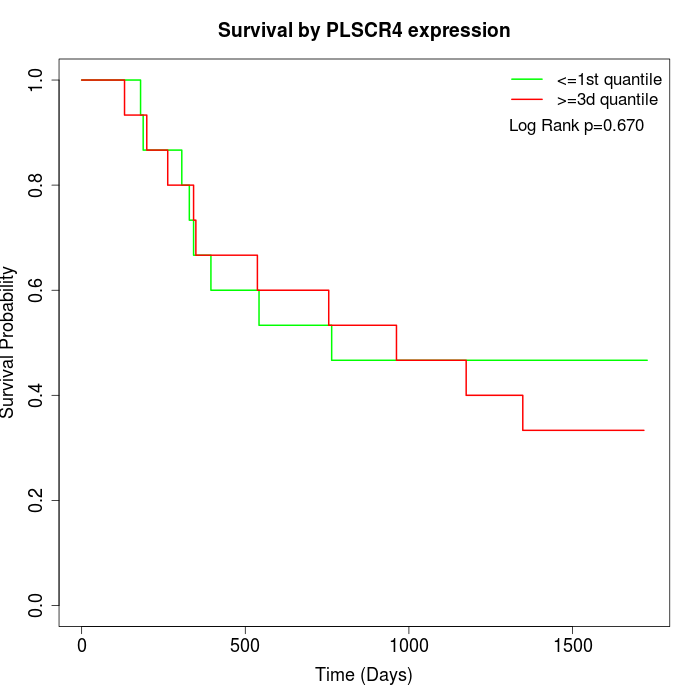
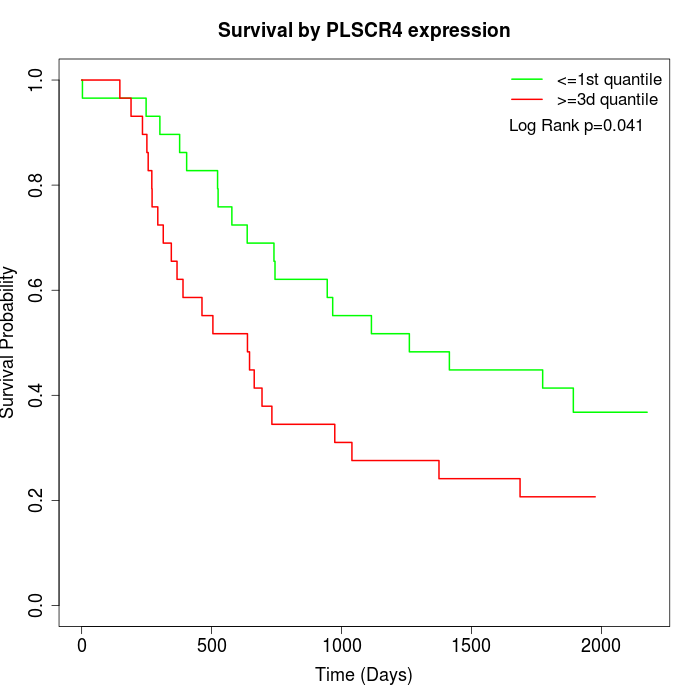
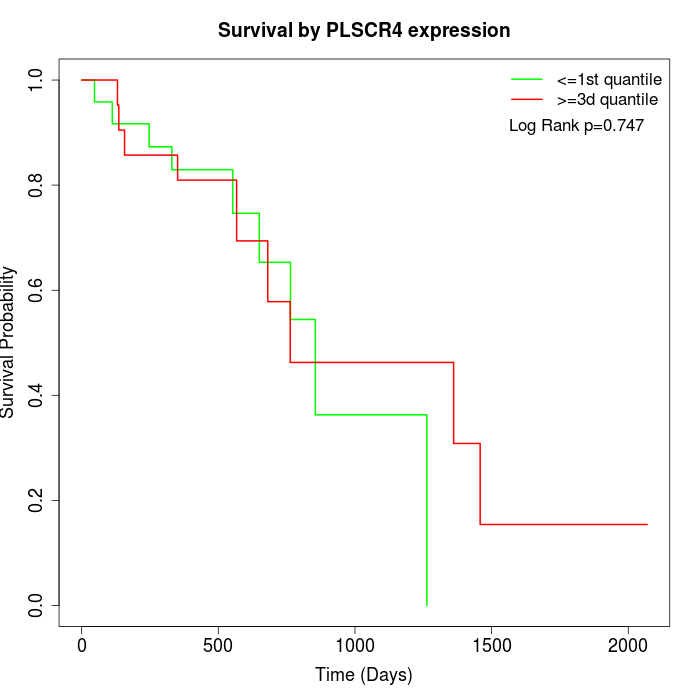
Survival by PLSCR4 expression:
Note: Click image to view full size file.
Copy number change of PLSCR4:
| Dataset | Gene | EntrezID | Gain | Loss | Normal | Detail |
|---|---|---|---|---|---|---|
| GSE15526 | PLSCR4 | 57088 | 22 | 0 | 8 | |
| GSE20123 | PLSCR4 | 57088 | 22 | 0 | 8 | |
| GSE43470 | PLSCR4 | 57088 | 23 | 0 | 20 | |
| GSE46452 | PLSCR4 | 57088 | 16 | 3 | 40 | |
| GSE47630 | PLSCR4 | 57088 | 19 | 3 | 18 | |
| GSE54993 | PLSCR4 | 57088 | 1 | 9 | 60 | |
| GSE54994 | PLSCR4 | 57088 | 34 | 1 | 18 | |
| GSE60625 | PLSCR4 | 57088 | 0 | 6 | 5 | |
| GSE74703 | PLSCR4 | 57088 | 20 | 0 | 16 | |
| GSE74704 | PLSCR4 | 57088 | 15 | 0 | 5 | |
| TCGA | PLSCR4 | 57088 | 66 | 2 | 28 |
Total number of gains: 238; Total number of losses: 24; Total Number of normals: 226.
Somatic mutations of PLSCR4:
Generating mutation plots.
Highly correlated genes for PLSCR4:
Showing top 20/606 corelated genes with mean PCC>0.5.
| Gene1 | Gene2 | Mean PCC | Num. Datasets | Num. PCC<0 | Num. PCC>0.5 |
|---|---|---|---|---|---|
| PLSCR4 | TMEM98 | 0.739639 | 3 | 0 | 3 |
| PLSCR4 | NDUFA8 | 0.721565 | 3 | 0 | 3 |
| PLSCR4 | SMAD5 | 0.720321 | 3 | 0 | 3 |
| PLSCR4 | TBC1D19 | 0.703173 | 3 | 0 | 3 |
| PLSCR4 | TNFSF12 | 0.701671 | 3 | 0 | 3 |
| PLSCR4 | TCEAL5 | 0.696211 | 3 | 0 | 3 |
| PLSCR4 | PIGC | 0.695413 | 3 | 0 | 3 |
| PLSCR4 | LRRN4CL | 0.691724 | 5 | 0 | 5 |
| PLSCR4 | ZZZ3 | 0.68905 | 4 | 0 | 4 |
| PLSCR4 | RAD21 | 0.686937 | 3 | 0 | 3 |
| PLSCR4 | RRN3 | 0.684146 | 3 | 0 | 3 |
| PLSCR4 | ZMAT1 | 0.673481 | 3 | 0 | 3 |
| PLSCR4 | NCBP2 | 0.673464 | 3 | 0 | 3 |
| PLSCR4 | ISCA1 | 0.67186 | 4 | 0 | 3 |
| PLSCR4 | ZC3H7B | 0.671676 | 3 | 0 | 3 |
| PLSCR4 | ZNF251 | 0.670909 | 3 | 0 | 3 |
| PLSCR4 | SNAPC5 | 0.668843 | 3 | 0 | 3 |
| PLSCR4 | MTX3 | 0.667996 | 4 | 0 | 4 |
| PLSCR4 | FRMD3 | 0.667898 | 4 | 0 | 4 |
| PLSCR4 | MYH10 | 0.666431 | 3 | 0 | 3 |
For details and further investigation, click here