| Full name: protein arginine methyltransferase 1 | Alias Symbol: HCP1|ANM1 | ||
| Type: protein-coding gene | Cytoband: 19q13.33 | ||
| Entrez ID: 3276 | HGNC ID: HGNC:5187 | Ensembl Gene: ENSG00000126457 | OMIM ID: 602950 |
| Drug and gene relationship at DGIdb | |||
Screen Evidence:
| |||
PRMT1 involved pathways:
| KEGG pathway | Description | View |
|---|---|---|
| hsa04068 | FoxO signaling pathway | |
| hsa04922 | Glucagon signaling pathway |
Expression of PRMT1:
| Dataset | Gene | EntrezID | Probe | Log2FC | Adj.pValue | Expression |
|---|---|---|---|---|---|---|
| GSE17351 | PRMT1 | 3276 | 206445_s_at | 0.8809 | 0.0174 | |
| GSE20347 | PRMT1 | 3276 | 206445_s_at | 0.6966 | 0.0001 | |
| GSE23400 | PRMT1 | 3276 | 206445_s_at | 0.6815 | 0.0000 | |
| GSE26886 | PRMT1 | 3276 | 206445_s_at | 0.7618 | 0.0055 | |
| GSE29001 | PRMT1 | 3276 | 206445_s_at | 0.5468 | 0.2454 | |
| GSE38129 | PRMT1 | 3276 | 206445_s_at | 0.8195 | 0.0001 | |
| GSE45670 | PRMT1 | 3276 | 206445_s_at | 0.2882 | 0.1326 | |
| GSE53622 | PRMT1 | 3276 | 19041 | 0.6038 | 0.0000 | |
| GSE53624 | PRMT1 | 3276 | 19041 | 0.7608 | 0.0000 | |
| GSE63941 | PRMT1 | 3276 | 206445_s_at | 1.2849 | 0.0012 | |
| GSE77861 | PRMT1 | 3276 | 206445_s_at | 0.7180 | 0.0208 | |
| GSE97050 | PRMT1 | 3276 | A_23_P39088 | 0.4033 | 0.1363 | |
| SRP007169 | PRMT1 | 3276 | RNAseq | 0.8289 | 0.0089 | |
| SRP008496 | PRMT1 | 3276 | RNAseq | 0.7000 | 0.0001 | |
| SRP064894 | PRMT1 | 3276 | RNAseq | 0.7295 | 0.0029 | |
| SRP133303 | PRMT1 | 3276 | RNAseq | 0.6717 | 0.0008 | |
| SRP159526 | PRMT1 | 3276 | RNAseq | 0.8093 | 0.0060 | |
| SRP193095 | PRMT1 | 3276 | RNAseq | 0.5488 | 0.0000 | |
| SRP219564 | PRMT1 | 3276 | RNAseq | 0.4269 | 0.1094 | |
| TCGA | PRMT1 | 3276 | RNAseq | 0.2841 | 0.0000 |
Upregulated datasets: 1; Downregulated datasets: 0.
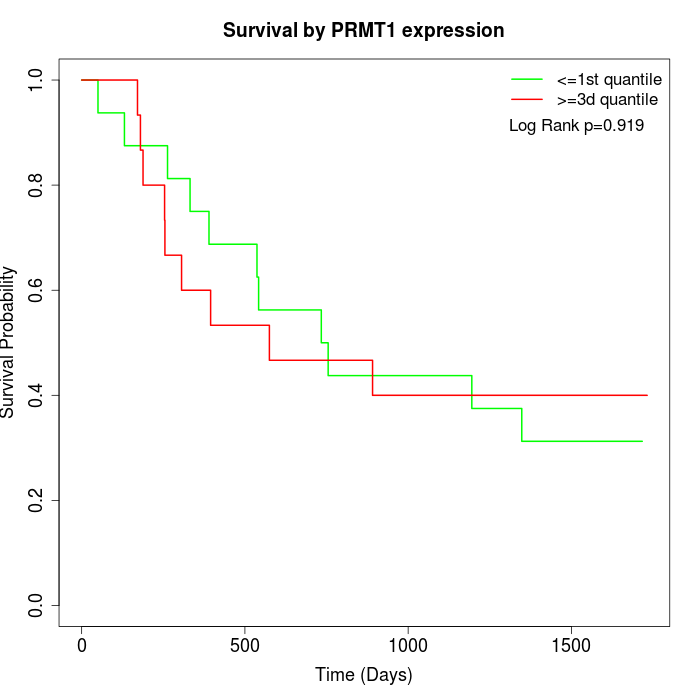
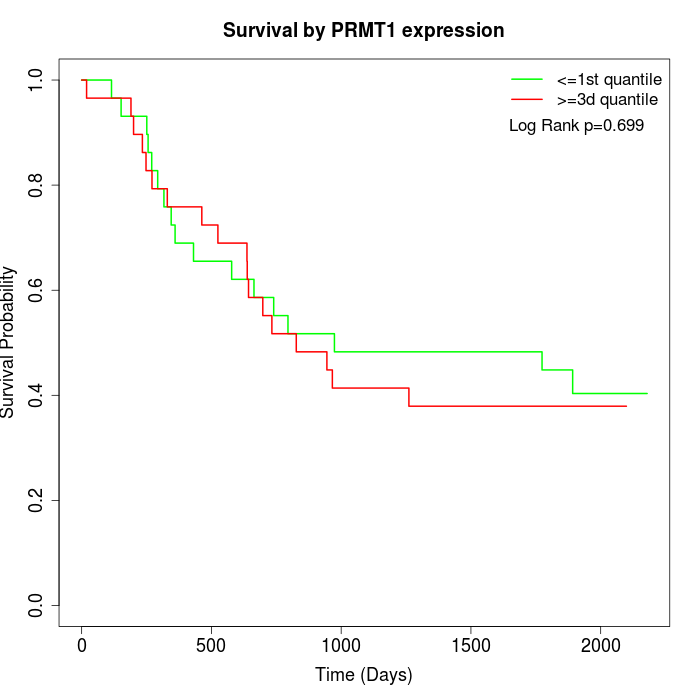
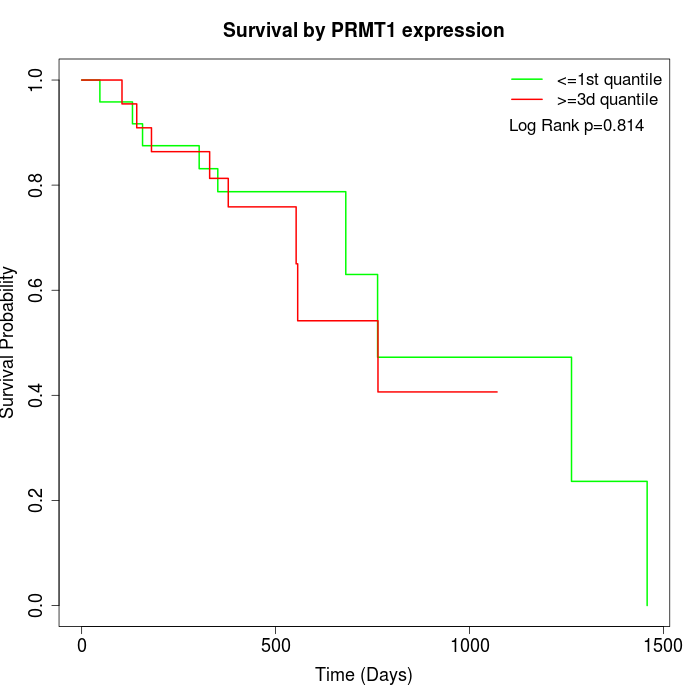
Survival by PRMT1 expression:
Note: Click image to view full size file.
Copy number change of PRMT1:
| Dataset | Gene | EntrezID | Gain | Loss | Normal | Detail |
|---|---|---|---|---|---|---|
| GSE15526 | PRMT1 | 3276 | 4 | 4 | 22 | |
| GSE20123 | PRMT1 | 3276 | 4 | 3 | 23 | |
| GSE43470 | PRMT1 | 3276 | 5 | 11 | 27 | |
| GSE46452 | PRMT1 | 3276 | 45 | 1 | 13 | |
| GSE47630 | PRMT1 | 3276 | 10 | 6 | 24 | |
| GSE54993 | PRMT1 | 3276 | 17 | 4 | 49 | |
| GSE54994 | PRMT1 | 3276 | 4 | 14 | 35 | |
| GSE60625 | PRMT1 | 3276 | 9 | 0 | 2 | |
| GSE74703 | PRMT1 | 3276 | 5 | 7 | 24 | |
| GSE74704 | PRMT1 | 3276 | 4 | 1 | 15 | |
| TCGA | PRMT1 | 3276 | 14 | 18 | 64 |
Total number of gains: 121; Total number of losses: 69; Total Number of normals: 298.
Somatic mutations of PRMT1:
Generating mutation plots.
Highly correlated genes for PRMT1:
Showing top 20/1997 corelated genes with mean PCC>0.5.
| Gene1 | Gene2 | Mean PCC | Num. Datasets | Num. PCC<0 | Num. PCC>0.5 |
|---|---|---|---|---|---|
| PRMT1 | RAB34 | 0.815022 | 3 | 0 | 3 |
| PRMT1 | EXOC6 | 0.800411 | 3 | 0 | 3 |
| PRMT1 | TRAPPC1 | 0.790186 | 3 | 0 | 3 |
| PRMT1 | PACS1 | 0.787343 | 3 | 0 | 3 |
| PRMT1 | SLC39A3 | 0.785806 | 4 | 0 | 4 |
| PRMT1 | CCDC14 | 0.763379 | 3 | 0 | 3 |
| PRMT1 | TUBA1B | 0.749446 | 9 | 0 | 9 |
| PRMT1 | EBNA1BP2 | 0.74684 | 3 | 0 | 3 |
| PRMT1 | MARCKSL1 | 0.741341 | 10 | 0 | 10 |
| PRMT1 | MRPL3 | 0.737942 | 9 | 0 | 9 |
| PRMT1 | CCT4 | 0.73732 | 11 | 0 | 11 |
| PRMT1 | FADS1 | 0.737117 | 3 | 0 | 3 |
| PRMT1 | SAAL1 | 0.735749 | 6 | 0 | 5 |
| PRMT1 | NF1 | 0.735025 | 3 | 0 | 3 |
| PRMT1 | NOP56 | 0.733814 | 3 | 0 | 3 |
| PRMT1 | CDK4 | 0.733462 | 13 | 0 | 11 |
| PRMT1 | KTI12 | 0.733305 | 3 | 0 | 3 |
| PRMT1 | FUS | 0.726749 | 11 | 0 | 10 |
| PRMT1 | ANAPC1 | 0.724845 | 3 | 0 | 3 |
| PRMT1 | TRIM28 | 0.723745 | 11 | 0 | 10 |
For details and further investigation, click here