| Full name: sarcolipin | Alias Symbol: MGC12301|MGC125854|MGC125855 | ||
| Type: protein-coding gene | Cytoband: 11q22.3 | ||
| Entrez ID: 6588 | HGNC ID: HGNC:11089 | Ensembl Gene: ENSG00000170290 | OMIM ID: 602203 |
Expression of SLN:
| Dataset | Gene | EntrezID | Probe | Log2FC | Adj.pValue | Expression |
|---|---|---|---|---|---|---|
| GSE17351 | SLN | 6588 | 205374_at | -0.0683 | 0.9356 | |
| GSE20347 | SLN | 6588 | 205374_at | -0.0147 | 0.9182 | |
| GSE23400 | SLN | 6588 | 205374_at | -0.0988 | 0.2051 | |
| GSE26886 | SLN | 6588 | 205374_at | 0.0081 | 0.9514 | |
| GSE29001 | SLN | 6588 | 205374_at | -0.1139 | 0.5855 | |
| GSE38129 | SLN | 6588 | 205374_at | -0.4001 | 0.0608 | |
| GSE45670 | SLN | 6588 | 205374_at | -0.0247 | 0.8464 | |
| GSE53622 | SLN | 6588 | 16220 | 0.5374 | 0.0918 | |
| GSE53624 | SLN | 6588 | 16220 | 0.8446 | 0.0001 | |
| GSE63941 | SLN | 6588 | 205374_at | 0.1349 | 0.4048 | |
| GSE77861 | SLN | 6588 | 205374_at | 0.0265 | 0.8141 | |
| GSE97050 | SLN | 6588 | A_23_P150343 | -1.7161 | 0.2017 | |
| TCGA | SLN | 6588 | RNAseq | 0.3456 | 0.6508 |
Upregulated datasets: 0; Downregulated datasets: 0.
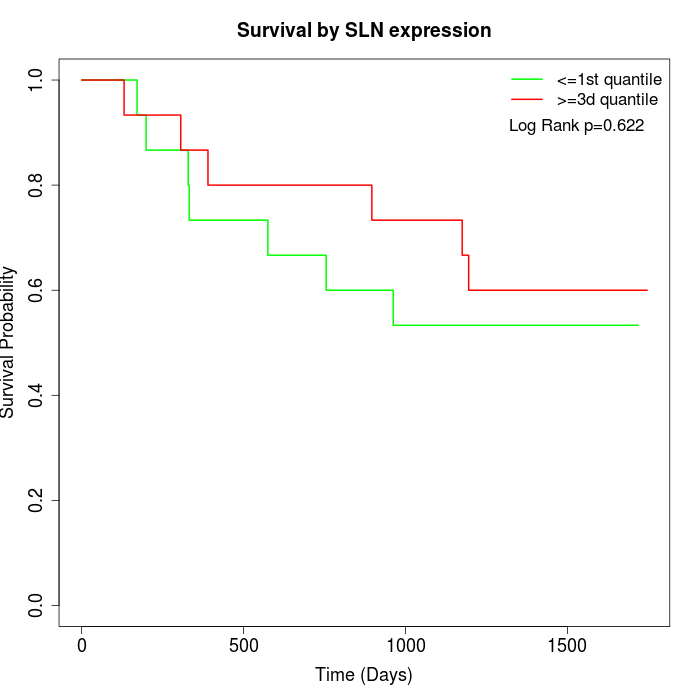
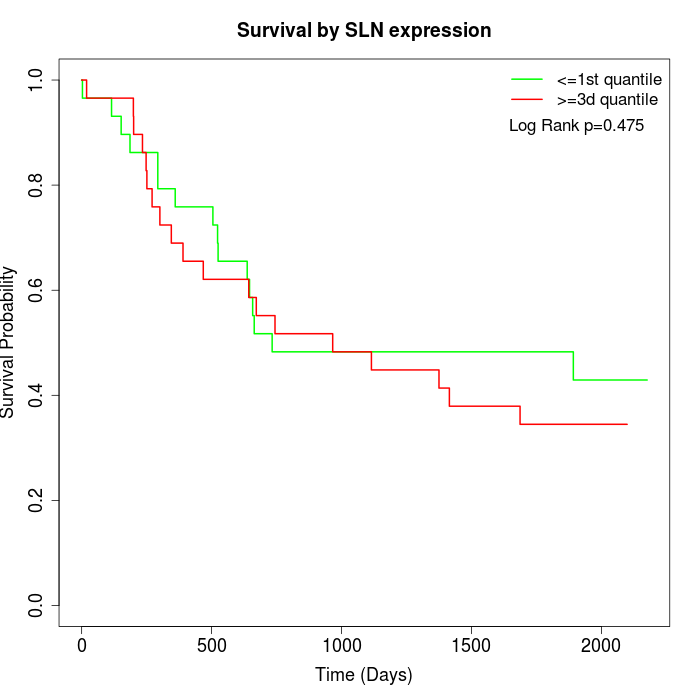
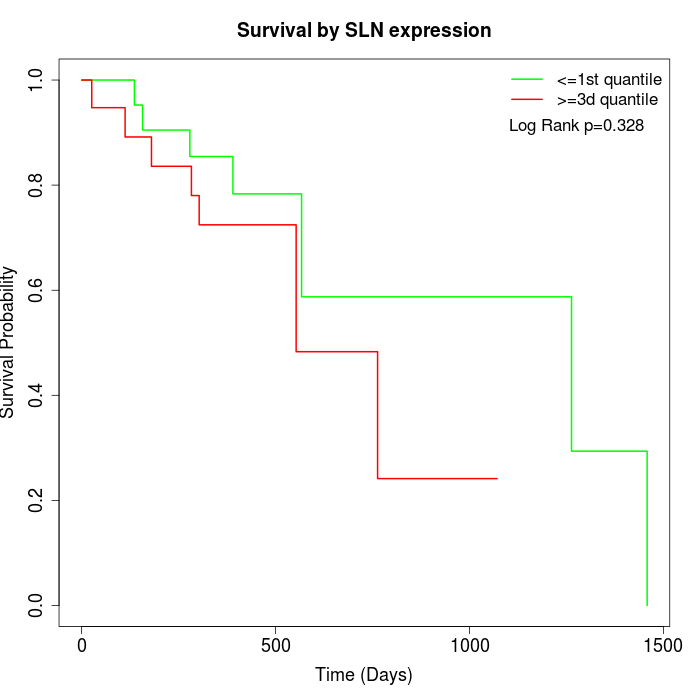
Survival by SLN expression:
Note: Click image to view full size file.
Copy number change of SLN:
| Dataset | Gene | EntrezID | Gain | Loss | Normal | Detail |
|---|---|---|---|---|---|---|
| GSE15526 | SLN | 6588 | 0 | 13 | 17 | |
| GSE20123 | SLN | 6588 | 0 | 13 | 17 | |
| GSE43470 | SLN | 6588 | 2 | 6 | 35 | |
| GSE46452 | SLN | 6588 | 3 | 26 | 30 | |
| GSE47630 | SLN | 6588 | 2 | 19 | 19 | |
| GSE54993 | SLN | 6588 | 9 | 1 | 60 | |
| GSE54994 | SLN | 6588 | 5 | 19 | 29 | |
| GSE60625 | SLN | 6588 | 0 | 3 | 8 | |
| GSE74703 | SLN | 6588 | 1 | 4 | 31 | |
| GSE74704 | SLN | 6588 | 0 | 9 | 11 | |
| TCGA | SLN | 6588 | 6 | 48 | 42 |
Total number of gains: 28; Total number of losses: 161; Total Number of normals: 299.
Somatic mutations of SLN:
Generating mutation plots.
Highly correlated genes for SLN:
Showing top 20/432 corelated genes with mean PCC>0.5.
| Gene1 | Gene2 | Mean PCC | Num. Datasets | Num. PCC<0 | Num. PCC>0.5 |
|---|---|---|---|---|---|
| SLN | CDC42EP2 | 0.745896 | 3 | 0 | 3 |
| SLN | KIAA1614 | 0.728208 | 3 | 0 | 3 |
| SLN | PCDH9 | 0.726627 | 3 | 0 | 3 |
| SLN | ABCA6 | 0.719186 | 4 | 0 | 4 |
| SLN | HABP4 | 0.711823 | 4 | 0 | 4 |
| SLN | STAC | 0.707069 | 3 | 0 | 3 |
| SLN | NKX6-1 | 0.705881 | 4 | 0 | 4 |
| SLN | TTN | 0.704447 | 3 | 0 | 3 |
| SLN | NACAD | 0.694591 | 4 | 0 | 4 |
| SLN | KCNC4 | 0.692094 | 3 | 0 | 3 |
| SLN | SLITRK3 | 0.68793 | 4 | 0 | 4 |
| SLN | TRIM45 | 0.683161 | 3 | 0 | 3 |
| SLN | TMEM8B | 0.682695 | 4 | 0 | 3 |
| SLN | NELL1 | 0.677772 | 3 | 0 | 3 |
| SLN | ZNF385D | 0.676846 | 4 | 0 | 4 |
| SLN | CNTNAP1 | 0.671474 | 4 | 0 | 4 |
| SLN | HAO2 | 0.67092 | 3 | 0 | 3 |
| SLN | MCAM | 0.669305 | 3 | 0 | 3 |
| SLN | INSRR | 0.668462 | 3 | 0 | 3 |
| SLN | CACNB2 | 0.667595 | 4 | 0 | 4 |
For details and further investigation, click here