| Full name: smoothened, frizzled class receptor | Alias Symbol: FZD11 | ||
| Type: protein-coding gene | Cytoband: 7q32.1 | ||
| Entrez ID: 6608 | HGNC ID: HGNC:11119 | Ensembl Gene: ENSG00000128602 | OMIM ID: 601500 |
| Related drugs: ARSENIC TRIOXIDE, BMS-833923, ERISMODEGIB, GEFITINIB, GLASDEGIB, LEQ506, PATIDEGIB, PHA-665752, POSACONAZOLE, SARIDEGIB... [more] | |||
SMO involved pathways:
| KEGG pathway | Description | View |
|---|---|---|
| hsa04340 | Hedgehog signaling pathway | |
| hsa04360 | Axon guidance | |
| hsa05200 | Pathways in cancer |
Expression of SMO:
| Dataset | Gene | EntrezID | Probe | Log2FC | Adj.pValue | Expression |
|---|---|---|---|---|---|---|
| GSE17351 | SMO | 6608 | 218629_at | -0.0490 | 0.8884 | |
| GSE20347 | SMO | 6608 | 218629_at | 0.0129 | 0.9247 | |
| GSE23400 | SMO | 6608 | 218629_at | -0.0436 | 0.3531 | |
| GSE26886 | SMO | 6608 | 218629_at | -0.4403 | 0.0080 | |
| GSE29001 | SMO | 6608 | 218629_at | -0.1173 | 0.5089 | |
| GSE38129 | SMO | 6608 | 218629_at | 0.0101 | 0.9412 | |
| GSE45670 | SMO | 6608 | 218629_at | -0.0069 | 0.9558 | |
| GSE53622 | SMO | 6608 | 54899 | 0.1162 | 0.3549 | |
| GSE53624 | SMO | 6608 | 54899 | 0.1048 | 0.3096 | |
| GSE63941 | SMO | 6608 | 218629_at | 0.1060 | 0.8102 | |
| GSE77861 | SMO | 6608 | 218629_at | -0.0235 | 0.8967 | |
| GSE97050 | SMO | 6608 | A_23_P70818 | -0.4092 | 0.1491 | |
| SRP007169 | SMO | 6608 | RNAseq | 0.1248 | 0.7644 | |
| SRP008496 | SMO | 6608 | RNAseq | 0.5141 | 0.0727 | |
| SRP064894 | SMO | 6608 | RNAseq | 0.3683 | 0.0473 | |
| SRP133303 | SMO | 6608 | RNAseq | 0.0022 | 0.9924 | |
| SRP159526 | SMO | 6608 | RNAseq | 0.8187 | 0.0100 | |
| SRP193095 | SMO | 6608 | RNAseq | -0.0353 | 0.8851 | |
| SRP219564 | SMO | 6608 | RNAseq | -0.0704 | 0.8036 | |
| TCGA | SMO | 6608 | RNAseq | 0.2223 | 0.1080 |
Upregulated datasets: 0; Downregulated datasets: 0.
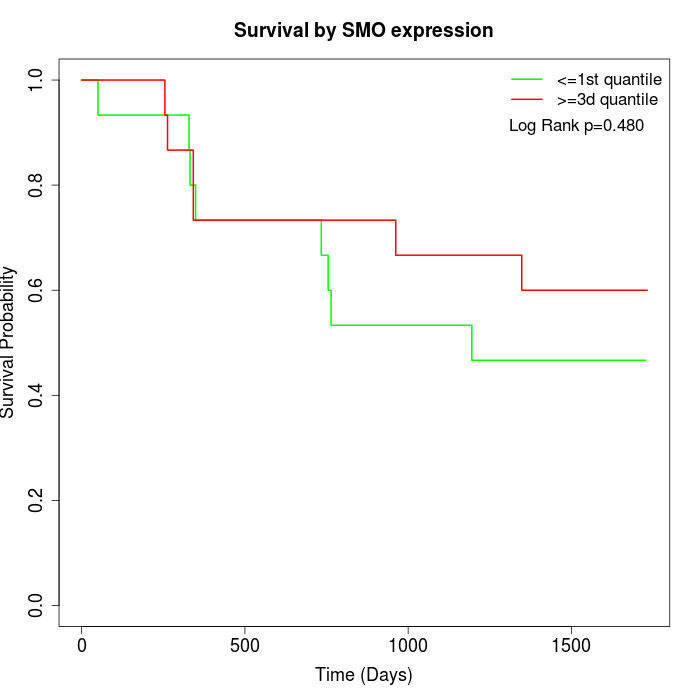
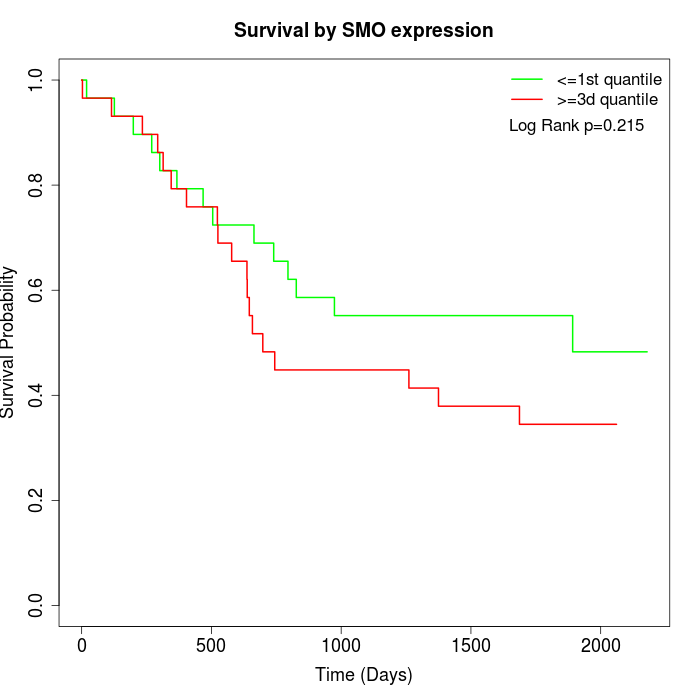
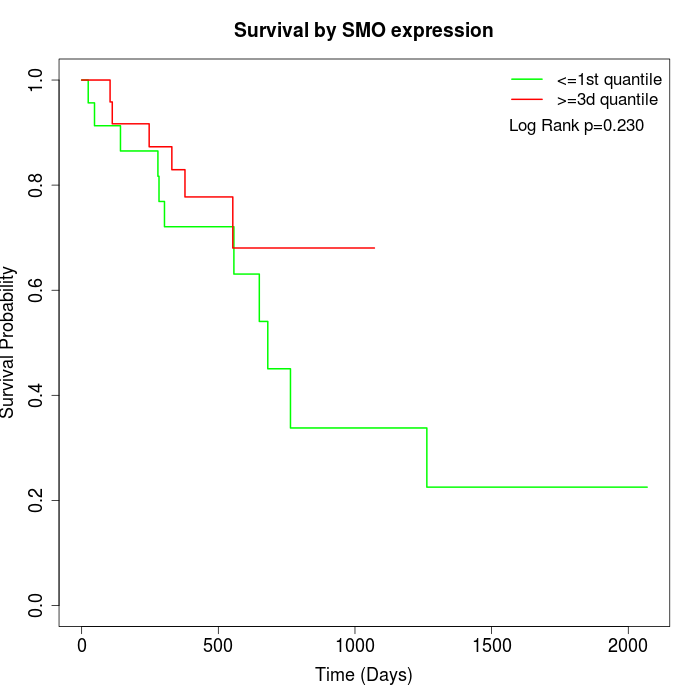
Survival by SMO expression:
Note: Click image to view full size file.
Copy number change of SMO:
| Dataset | Gene | EntrezID | Gain | Loss | Normal | Detail |
|---|---|---|---|---|---|---|
| GSE15526 | SMO | 6608 | 8 | 1 | 21 | |
| GSE20123 | SMO | 6608 | 8 | 1 | 21 | |
| GSE43470 | SMO | 6608 | 2 | 3 | 38 | |
| GSE46452 | SMO | 6608 | 7 | 2 | 50 | |
| GSE47630 | SMO | 6608 | 7 | 4 | 29 | |
| GSE54993 | SMO | 6608 | 3 | 5 | 62 | |
| GSE54994 | SMO | 6608 | 8 | 7 | 38 | |
| GSE60625 | SMO | 6608 | 0 | 0 | 11 | |
| GSE74703 | SMO | 6608 | 2 | 3 | 31 | |
| GSE74704 | SMO | 6608 | 4 | 1 | 15 | |
| TCGA | SMO | 6608 | 29 | 21 | 46 |
Total number of gains: 78; Total number of losses: 48; Total Number of normals: 362.
Somatic mutations of SMO:
Generating mutation plots.
Highly correlated genes for SMO:
Showing top 20/284 corelated genes with mean PCC>0.5.
| Gene1 | Gene2 | Mean PCC | Num. Datasets | Num. PCC<0 | Num. PCC>0.5 |
|---|---|---|---|---|---|
| SMO | STAT6 | 0.760313 | 3 | 0 | 3 |
| SMO | ZDHHC3 | 0.747938 | 3 | 0 | 3 |
| SMO | ADCY6 | 0.738971 | 3 | 0 | 3 |
| SMO | ZNF467 | 0.726655 | 4 | 0 | 3 |
| SMO | ACO2 | 0.718417 | 3 | 0 | 3 |
| SMO | BBS1 | 0.710533 | 3 | 0 | 3 |
| SMO | ARRB1 | 0.709654 | 3 | 0 | 3 |
| SMO | SERPINA5 | 0.708868 | 4 | 0 | 3 |
| SMO | COBLL1 | 0.705134 | 3 | 0 | 3 |
| SMO | CELA3A | 0.702093 | 3 | 0 | 3 |
| SMO | RAB5B | 0.700933 | 3 | 0 | 3 |
| SMO | HDAC11 | 0.697081 | 4 | 0 | 3 |
| SMO | GPATCH8 | 0.688221 | 3 | 0 | 3 |
| SMO | AKAP7 | 0.687114 | 3 | 0 | 3 |
| SMO | RBP2 | 0.683066 | 3 | 0 | 3 |
| SMO | CBL | 0.67807 | 4 | 0 | 3 |
| SMO | MTA3 | 0.676296 | 3 | 0 | 3 |
| SMO | CHRNE | 0.674355 | 3 | 0 | 3 |
| SMO | ZNF516 | 0.667324 | 4 | 0 | 3 |
| SMO | TMEM214 | 0.667087 | 4 | 0 | 3 |
For details and further investigation, click here