| Full name: TCL1 upstream neural differentiation-associated RNA | Alias Symbol: TUNA|HI-LNC78 | ||
| Type: non-coding RNA | Cytoband: 14q32.2 | ||
| Entrez ID: 100507043 | HGNC ID: HGNC:44088 | Ensembl Gene: ENSG00000250366 | OMIM ID: 615719 |
Expression of TUNAR:
| Dataset | Gene | EntrezID | Probe | Log2FC | Adj.pValue | Expression |
|---|---|---|---|---|---|---|
| GSE17351 | TUNAR | 100507043 | 232111_at | -0.3156 | 0.2777 | |
| GSE26886 | TUNAR | 100507043 | 232111_at | -0.3211 | 0.0030 | |
| GSE45670 | TUNAR | 100507043 | 232111_at | -0.0675 | 0.6610 | |
| GSE53622 | TUNAR | 100507043 | 86562 | -0.4687 | 0.0020 | |
| GSE53624 | TUNAR | 100507043 | 86562 | -0.5960 | 0.0001 | |
| GSE63941 | TUNAR | 100507043 | 232111_at | -0.1179 | 0.4242 | |
| GSE77861 | TUNAR | 100507043 | 232111_at | -0.0825 | 0.3849 | |
| SRP133303 | TUNAR | 100507043 | RNAseq | 0.0831 | 0.5634 | |
| SRP193095 | TUNAR | 100507043 | RNAseq | 0.2751 | 0.0728 |
Upregulated datasets: 0; Downregulated datasets: 0.
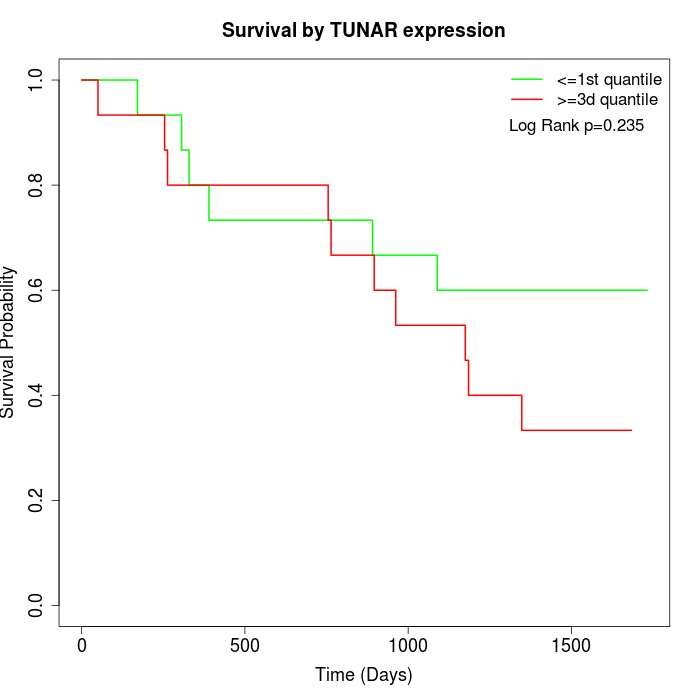
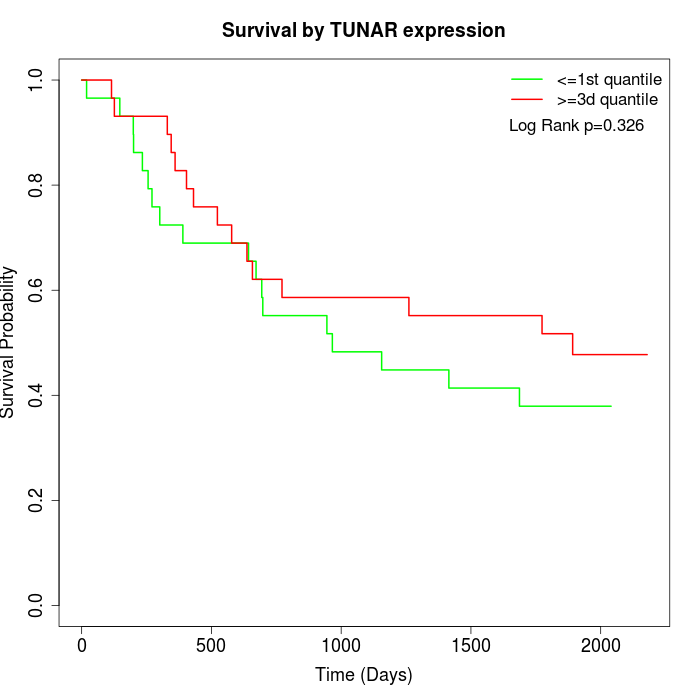
Survival by TUNAR expression:
Note: Click image to view full size file.
Copy number change of TUNAR:
| Dataset | Gene | EntrezID | Gain | Loss | Normal | Detail |
|---|---|---|---|---|---|---|
| GSE15526 | TUNAR | 100507043 | 10 | 4 | 16 | |
| GSE20123 | TUNAR | 100507043 | 10 | 3 | 17 | |
| GSE43470 | TUNAR | 100507043 | 8 | 3 | 32 | |
| GSE46452 | TUNAR | 100507043 | 16 | 3 | 40 | |
| GSE47630 | TUNAR | 100507043 | 11 | 8 | 21 | |
| GSE54993 | TUNAR | 100507043 | 3 | 8 | 59 | |
| GSE54994 | TUNAR | 100507043 | 21 | 4 | 28 | |
| GSE60625 | TUNAR | 100507043 | 0 | 2 | 9 | |
| GSE74703 | TUNAR | 100507043 | 6 | 3 | 27 | |
| GSE74704 | TUNAR | 100507043 | 6 | 3 | 11 |
Total number of gains: 91; Total number of losses: 41; Total Number of normals: 260.
Somatic mutations of TUNAR:
Generating mutation plots.
Highly correlated genes for TUNAR:
Showing top 20/135 corelated genes with mean PCC>0.5.
| Gene1 | Gene2 | Mean PCC | Num. Datasets | Num. PCC<0 | Num. PCC>0.5 |
|---|---|---|---|---|---|
| TUNAR | GOT1 | 0.700425 | 3 | 0 | 3 |
| TUNAR | PCYOX1 | 0.69721 | 3 | 0 | 3 |
| TUNAR | UGP2 | 0.688712 | 3 | 0 | 3 |
| TUNAR | DYNLT3 | 0.686522 | 3 | 0 | 3 |
| TUNAR | DLGAP1-AS3 | 0.681571 | 3 | 0 | 3 |
| TUNAR | GHITM | 0.678209 | 3 | 0 | 3 |
| TUNAR | CRYBG3 | 0.657933 | 3 | 0 | 3 |
| TUNAR | PPP1R7 | 0.65545 | 3 | 0 | 3 |
| TUNAR | PRADC1 | 0.651447 | 3 | 0 | 3 |
| TUNAR | CMPK1 | 0.649406 | 4 | 0 | 4 |
| TUNAR | SPAG17 | 0.649048 | 3 | 0 | 3 |
| TUNAR | FAM71C | 0.648263 | 4 | 0 | 3 |
| TUNAR | MFSD5 | 0.640688 | 3 | 0 | 3 |
| TUNAR | CLPX | 0.638346 | 3 | 0 | 3 |
| TUNAR | CYP2C18 | 0.636916 | 3 | 0 | 3 |
| TUNAR | RHOV | 0.632607 | 4 | 0 | 3 |
| TUNAR | ORMDL2 | 0.630021 | 3 | 0 | 3 |
| TUNAR | RNF11 | 0.629298 | 4 | 0 | 3 |
| TUNAR | CHMP2A | 0.626738 | 4 | 0 | 3 |
| TUNAR | CHP1 | 0.626561 | 4 | 0 | 3 |
For details and further investigation, click here