| Full name: alpha and gamma adaptin binding protein | Alias Symbol: FLJ11506|p34 | ||
| Type: protein-coding gene | Cytoband: 15q23 | ||
| Entrez ID: 79719 | HGNC ID: HGNC:25662 | Ensembl Gene: ENSG00000103591 | OMIM ID: 614888 |
Expression of AAGAB:
| Dataset | Gene | EntrezID | Probe | Log2FC | Adj.pValue | Expression |
|---|---|---|---|---|---|---|
| GSE17351 | AAGAB | 79719 | 202852_s_at | 0.1905 | 0.7975 | |
| GSE20347 | AAGAB | 79719 | 202852_s_at | 0.1399 | 0.5016 | |
| GSE23400 | AAGAB | 79719 | 202852_s_at | 0.2241 | 0.0024 | |
| GSE26886 | AAGAB | 79719 | 202852_s_at | 0.2447 | 0.4210 | |
| GSE29001 | AAGAB | 79719 | 202852_s_at | 0.1075 | 0.7520 | |
| GSE38129 | AAGAB | 79719 | 202852_s_at | 0.3597 | 0.0299 | |
| GSE45670 | AAGAB | 79719 | 202852_s_at | 0.5365 | 0.0001 | |
| GSE53622 | AAGAB | 79719 | 46624 | 0.2849 | 0.0008 | |
| GSE53624 | AAGAB | 79719 | 46624 | 0.2034 | 0.0001 | |
| GSE63941 | AAGAB | 79719 | 202852_s_at | 1.4466 | 0.0005 | |
| GSE77861 | AAGAB | 79719 | 202852_s_at | 0.3767 | 0.1456 | |
| GSE97050 | AAGAB | 79719 | A_23_P37545 | 0.3785 | 0.2027 | |
| SRP007169 | AAGAB | 79719 | RNAseq | -0.2720 | 0.4469 | |
| SRP008496 | AAGAB | 79719 | RNAseq | -0.1686 | 0.4346 | |
| SRP064894 | AAGAB | 79719 | RNAseq | 0.0567 | 0.6669 | |
| SRP133303 | AAGAB | 79719 | RNAseq | 0.4984 | 0.0050 | |
| SRP159526 | AAGAB | 79719 | RNAseq | -0.0208 | 0.9565 | |
| SRP193095 | AAGAB | 79719 | RNAseq | -0.0661 | 0.5028 | |
| SRP219564 | AAGAB | 79719 | RNAseq | -0.0603 | 0.7503 | |
| TCGA | AAGAB | 79719 | RNAseq | -0.0002 | 0.9969 |
Upregulated datasets: 1; Downregulated datasets: 0.
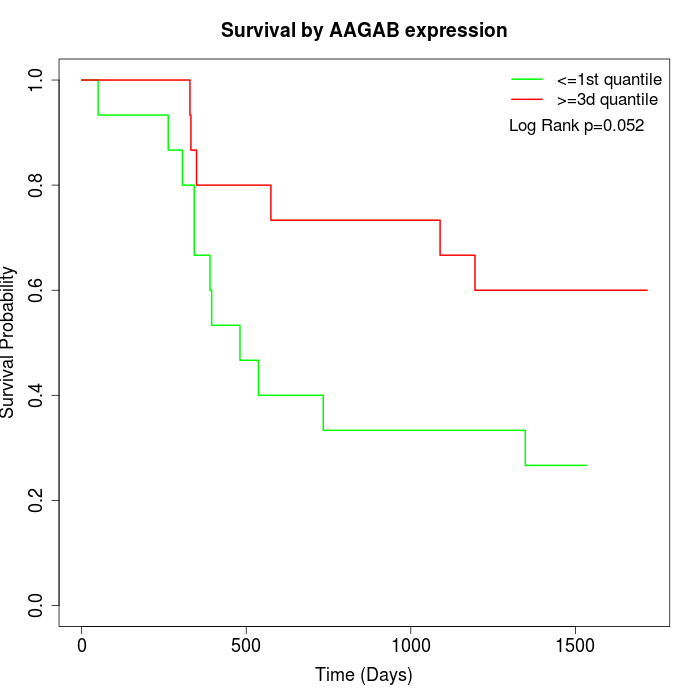
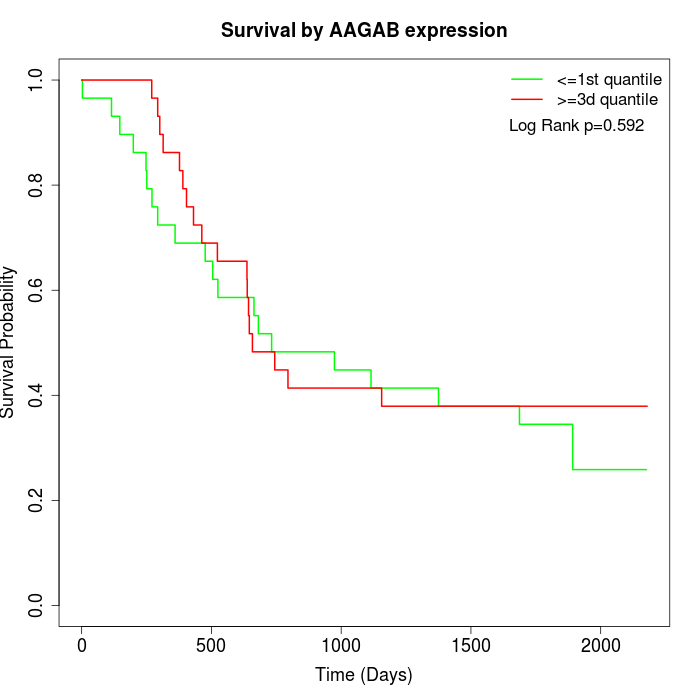
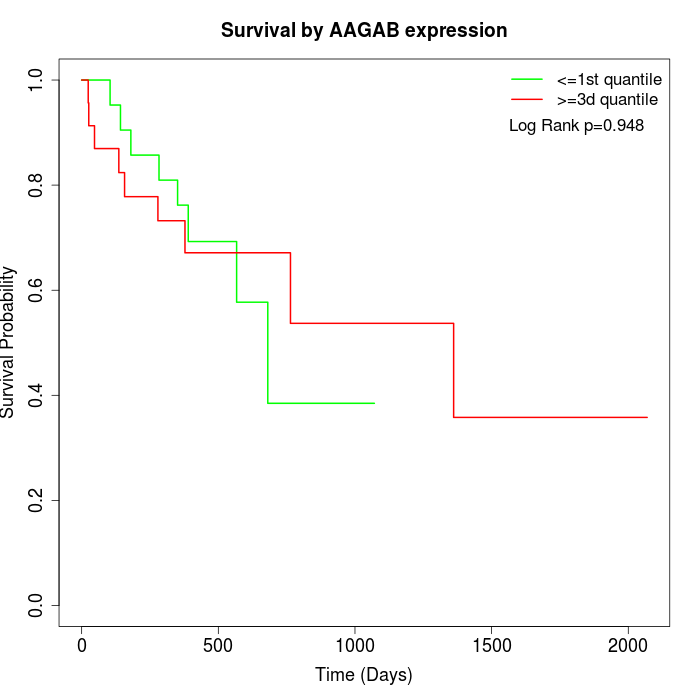
Survival by AAGAB expression:
Note: Click image to view full size file.
Copy number change of AAGAB:
| Dataset | Gene | EntrezID | Gain | Loss | Normal | Detail |
|---|---|---|---|---|---|---|
| GSE15526 | AAGAB | 79719 | 8 | 2 | 20 | |
| GSE20123 | AAGAB | 79719 | 8 | 2 | 20 | |
| GSE43470 | AAGAB | 79719 | 4 | 6 | 33 | |
| GSE46452 | AAGAB | 79719 | 3 | 7 | 49 | |
| GSE47630 | AAGAB | 79719 | 8 | 10 | 22 | |
| GSE54993 | AAGAB | 79719 | 5 | 6 | 59 | |
| GSE54994 | AAGAB | 79719 | 6 | 7 | 40 | |
| GSE60625 | AAGAB | 79719 | 4 | 0 | 7 | |
| GSE74703 | AAGAB | 79719 | 4 | 3 | 29 | |
| GSE74704 | AAGAB | 79719 | 3 | 2 | 15 | |
| TCGA | AAGAB | 79719 | 10 | 15 | 71 |
Total number of gains: 63; Total number of losses: 60; Total Number of normals: 365.
Somatic mutations of AAGAB:
Generating mutation plots.
Highly correlated genes for AAGAB:
Showing top 20/588 corelated genes with mean PCC>0.5.
| Gene1 | Gene2 | Mean PCC | Num. Datasets | Num. PCC<0 | Num. PCC>0.5 |
|---|---|---|---|---|---|
| AAGAB | ITGB8 | 0.815185 | 3 | 0 | 3 |
| AAGAB | PLS1 | 0.789626 | 3 | 0 | 3 |
| AAGAB | IREB2 | 0.7771 | 3 | 0 | 3 |
| AAGAB | AFTPH | 0.773744 | 3 | 0 | 3 |
| AAGAB | NPRL3 | 0.763778 | 3 | 0 | 3 |
| AAGAB | MRPL38 | 0.755584 | 3 | 0 | 3 |
| AAGAB | ALG10 | 0.752898 | 3 | 0 | 3 |
| AAGAB | ARHGEF5 | 0.735612 | 4 | 0 | 4 |
| AAGAB | HMGB3 | 0.727364 | 3 | 0 | 3 |
| AAGAB | NDUFV1 | 0.723782 | 3 | 0 | 3 |
| AAGAB | PLEKHA8 | 0.722177 | 3 | 0 | 3 |
| AAGAB | PPP1R9B | 0.709925 | 3 | 0 | 3 |
| AAGAB | CNOT1 | 0.70872 | 3 | 0 | 3 |
| AAGAB | EEFSEC | 0.70657 | 3 | 0 | 3 |
| AAGAB | SUPT7L | 0.706222 | 3 | 0 | 3 |
| AAGAB | YWHAE | 0.685366 | 4 | 0 | 4 |
| AAGAB | COCH | 0.681803 | 5 | 0 | 5 |
| AAGAB | ENTPD7 | 0.680706 | 4 | 0 | 3 |
| AAGAB | MREG | 0.680134 | 4 | 0 | 3 |
| AAGAB | PIGF | 0.678478 | 4 | 0 | 4 |
For details and further investigation, click here