| Full name: BRMS1 transcriptional repressor and anoikis regulator | Alias Symbol: DKFZP564A063 | ||
| Type: protein-coding gene | Cytoband: 11q13.2 | ||
| Entrez ID: 25855 | HGNC ID: HGNC:17262 | Ensembl Gene: ENSG00000174744 | OMIM ID: 606259 |
Screen Evidence:
| |||
Expression of BRMS1:
| Dataset | Gene | EntrezID | Probe | Log2FC | Adj.pValue | Expression |
|---|---|---|---|---|---|---|
| GSE17351 | BRMS1 | 25855 | 215631_s_at | 0.8842 | 0.0494 | |
| GSE20347 | BRMS1 | 25855 | 215631_s_at | 0.6795 | 0.0002 | |
| GSE23400 | BRMS1 | 25855 | 215631_s_at | 0.3692 | 0.0000 | |
| GSE26886 | BRMS1 | 25855 | 215631_s_at | 0.1032 | 0.7158 | |
| GSE29001 | BRMS1 | 25855 | 215631_s_at | 0.4682 | 0.0190 | |
| GSE38129 | BRMS1 | 25855 | 215631_s_at | 0.7070 | 0.0000 | |
| GSE45670 | BRMS1 | 25855 | 215631_s_at | 0.5980 | 0.0001 | |
| GSE53622 | BRMS1 | 25855 | 37788 | 0.7506 | 0.0000 | |
| GSE53624 | BRMS1 | 25855 | 37788 | 0.7686 | 0.0000 | |
| GSE63941 | BRMS1 | 25855 | 215631_s_at | 0.7845 | 0.0378 | |
| GSE77861 | BRMS1 | 25855 | 215631_s_at | 0.5363 | 0.0031 | |
| GSE97050 | BRMS1 | 25855 | A_33_P3367017 | 0.6755 | 0.2589 | |
| SRP007169 | BRMS1 | 25855 | RNAseq | 1.2512 | 0.0236 | |
| SRP008496 | BRMS1 | 25855 | RNAseq | 1.5001 | 0.0012 | |
| SRP064894 | BRMS1 | 25855 | RNAseq | 0.6984 | 0.0103 | |
| SRP133303 | BRMS1 | 25855 | RNAseq | 0.9314 | 0.0000 | |
| SRP159526 | BRMS1 | 25855 | RNAseq | 0.5182 | 0.0239 | |
| SRP193095 | BRMS1 | 25855 | RNAseq | 0.4887 | 0.0001 | |
| SRP219564 | BRMS1 | 25855 | RNAseq | 0.8324 | 0.0379 | |
| TCGA | BRMS1 | 25855 | RNAseq | 0.2793 | 0.0000 |
Upregulated datasets: 2; Downregulated datasets: 0.
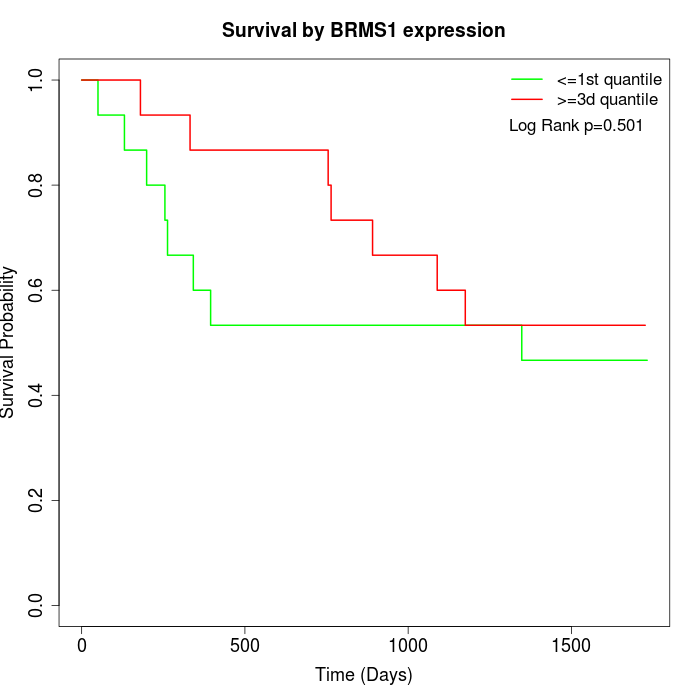
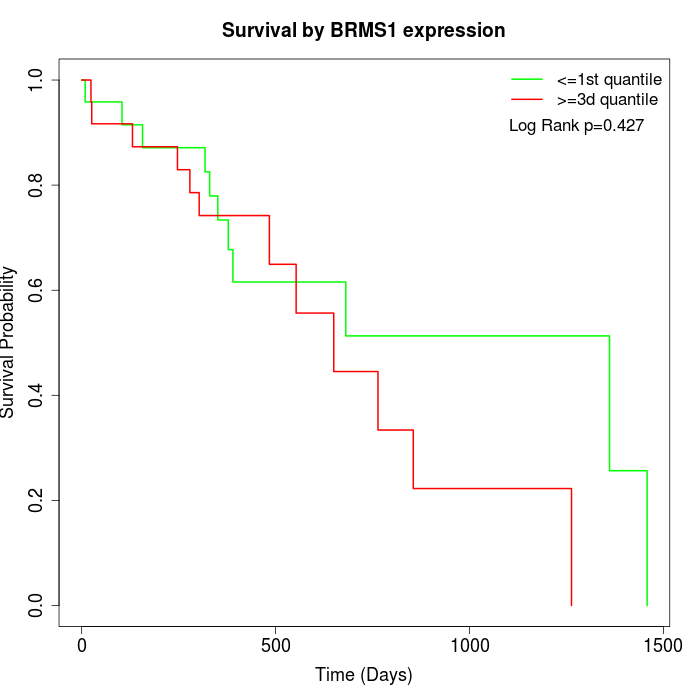
Survival by BRMS1 expression:
Note: Click image to view full size file.
Copy number change of BRMS1:
| Dataset | Gene | EntrezID | Gain | Loss | Normal | Detail |
|---|---|---|---|---|---|---|
| GSE15526 | BRMS1 | 25855 | 9 | 4 | 17 | |
| GSE20123 | BRMS1 | 25855 | 9 | 4 | 17 | |
| GSE43470 | BRMS1 | 25855 | 6 | 1 | 36 | |
| GSE46452 | BRMS1 | 25855 | 11 | 3 | 45 | |
| GSE47630 | BRMS1 | 25855 | 8 | 5 | 27 | |
| GSE54993 | BRMS1 | 25855 | 3 | 0 | 67 | |
| GSE54994 | BRMS1 | 25855 | 8 | 5 | 40 | |
| GSE60625 | BRMS1 | 25855 | 0 | 3 | 8 | |
| GSE74703 | BRMS1 | 25855 | 4 | 0 | 32 | |
| GSE74704 | BRMS1 | 25855 | 7 | 2 | 11 | |
| TCGA | BRMS1 | 25855 | 25 | 6 | 65 |
Total number of gains: 90; Total number of losses: 33; Total Number of normals: 365.
Somatic mutations of BRMS1:
Generating mutation plots.
Highly correlated genes for BRMS1:
Showing top 20/1547 corelated genes with mean PCC>0.5.
| Gene1 | Gene2 | Mean PCC | Num. Datasets | Num. PCC<0 | Num. PCC>0.5 |
|---|---|---|---|---|---|
| BRMS1 | PSMG3 | 0.783489 | 6 | 0 | 6 |
| BRMS1 | DPP3 | 0.750021 | 11 | 0 | 11 |
| BRMS1 | CCDC86 | 0.747957 | 7 | 0 | 7 |
| BRMS1 | ADRM1 | 0.745524 | 10 | 0 | 10 |
| BRMS1 | CFL1 | 0.740137 | 10 | 0 | 10 |
| BRMS1 | MICALCL | 0.727196 | 3 | 0 | 3 |
| BRMS1 | GEN1 | 0.726174 | 4 | 0 | 4 |
| BRMS1 | MRPL14 | 0.723365 | 5 | 0 | 5 |
| BRMS1 | SOGA1 | 0.722989 | 3 | 0 | 3 |
| BRMS1 | ZNF114 | 0.721201 | 3 | 0 | 3 |
| BRMS1 | YIF1A | 0.720604 | 13 | 0 | 13 |
| BRMS1 | UHRF1 | 0.719862 | 6 | 0 | 6 |
| BRMS1 | UBE2C | 0.717435 | 11 | 0 | 10 |
| BRMS1 | RRP36 | 0.712664 | 5 | 0 | 5 |
| BRMS1 | EME1 | 0.712133 | 5 | 0 | 4 |
| BRMS1 | BANF1 | 0.710326 | 11 | 0 | 10 |
| BRMS1 | RAD18 | 0.707589 | 3 | 0 | 3 |
| BRMS1 | STIP1 | 0.70599 | 12 | 0 | 11 |
| BRMS1 | WDR74 | 0.703841 | 11 | 0 | 10 |
| BRMS1 | RCC2 | 0.702395 | 6 | 0 | 6 |
For details and further investigation, click here