| Full name: checkpoint kinase 2 | Alias Symbol: CDS1|CHK2|HuCds1|PP1425|bA444G7 | ||
| Type: protein-coding gene | Cytoband: 22q12.1 | ||
| Entrez ID: 11200 | HGNC ID: HGNC:16627 | Ensembl Gene: ENSG00000183765 | OMIM ID: 604373 |
| Related drugs: AZD-7762, CHEMBL1236539, CHEMBL179583, ISOGRANULATIMIDE, OLAPARIB, PF-00477736, PREXASERTIB, XL-844... [more] | |||
Screen Evidence:
| |||
CHEK2 involved pathways:
| KEGG pathway | Description | View |
|---|---|---|
| hsa04110 | Cell cycle | |
| hsa04115 | p53 signaling pathway | |
| hsa05166 | HTLV-I infection |
Expression of CHEK2:
| Dataset | Gene | EntrezID | Probe | Log2FC | Adj.pValue | Expression |
|---|---|---|---|---|---|---|
| GSE17351 | CHEK2 | 11200 | 210416_s_at | 0.8316 | 0.0793 | |
| GSE20347 | CHEK2 | 11200 | 210416_s_at | 0.9583 | 0.0000 | |
| GSE23400 | CHEK2 | 11200 | 210416_s_at | 0.4773 | 0.0000 | |
| GSE26886 | CHEK2 | 11200 | 210416_s_at | 0.2960 | 0.1273 | |
| GSE29001 | CHEK2 | 11200 | 210416_s_at | 0.7977 | 0.0006 | |
| GSE38129 | CHEK2 | 11200 | 210416_s_at | 1.0876 | 0.0000 | |
| GSE45670 | CHEK2 | 11200 | 210416_s_at | 0.9393 | 0.0000 | |
| GSE53622 | CHEK2 | 11200 | 8948 | 0.7900 | 0.0000 | |
| GSE53624 | CHEK2 | 11200 | 25756 | 1.0398 | 0.0000 | |
| GSE63941 | CHEK2 | 11200 | 210416_s_at | 0.4887 | 0.4498 | |
| GSE77861 | CHEK2 | 11200 | 210416_s_at | 0.6727 | 0.0495 | |
| GSE97050 | CHEK2 | 11200 | A_33_P3386760 | 0.6238 | 0.1657 | |
| SRP007169 | CHEK2 | 11200 | RNAseq | 1.0031 | 0.0128 | |
| SRP008496 | CHEK2 | 11200 | RNAseq | 0.7847 | 0.0148 | |
| SRP064894 | CHEK2 | 11200 | RNAseq | 1.0226 | 0.0000 | |
| SRP133303 | CHEK2 | 11200 | RNAseq | 0.6399 | 0.0069 | |
| SRP159526 | CHEK2 | 11200 | RNAseq | 1.1792 | 0.0000 | |
| SRP193095 | CHEK2 | 11200 | RNAseq | 0.6225 | 0.0000 | |
| SRP219564 | CHEK2 | 11200 | RNAseq | 1.0269 | 0.0036 | |
| TCGA | CHEK2 | 11200 | RNAseq | 0.8624 | 0.0000 |
Upregulated datasets: 6; Downregulated datasets: 0.
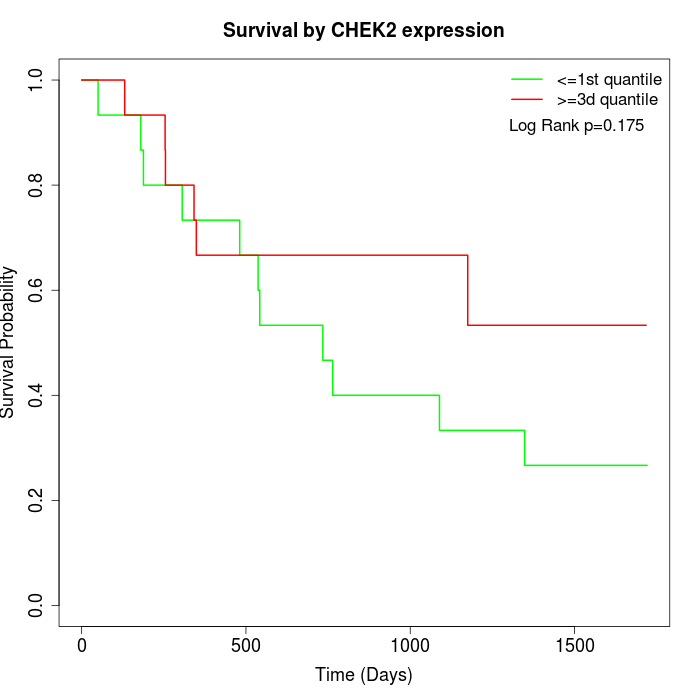
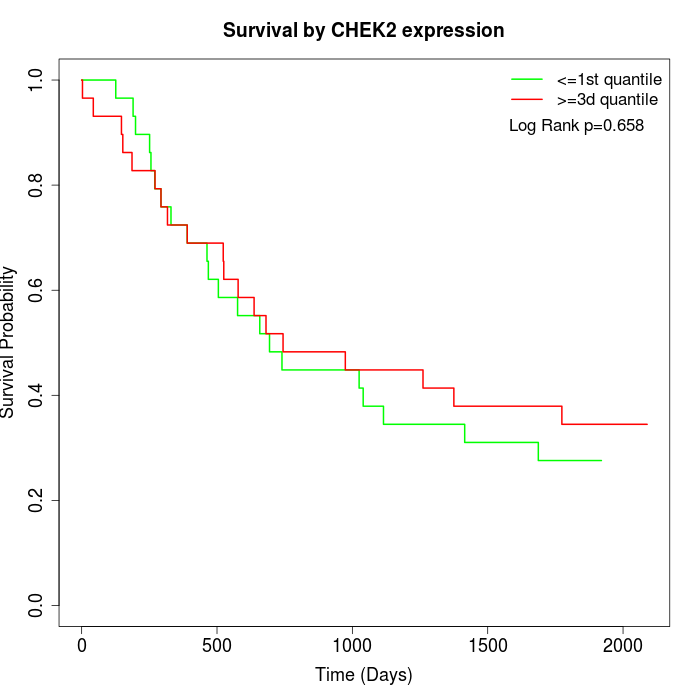
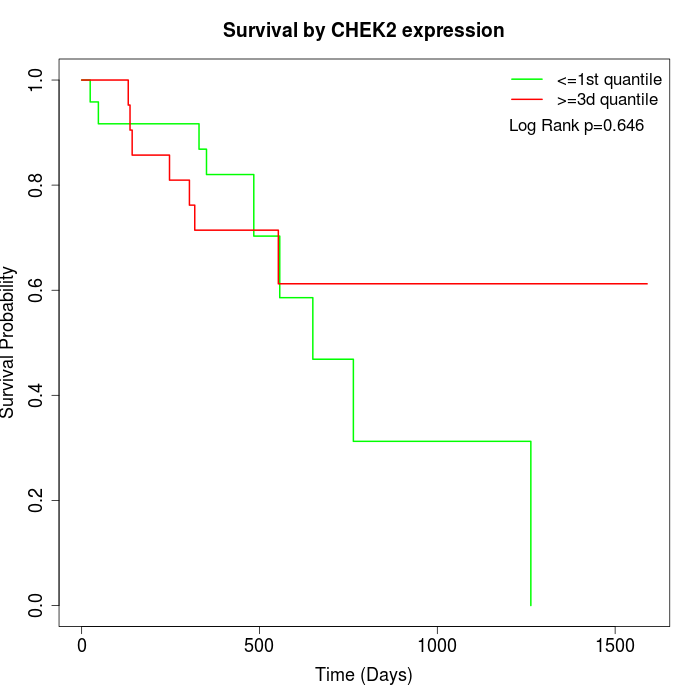
Survival by CHEK2 expression:
Note: Click image to view full size file.
Copy number change of CHEK2:
| Dataset | Gene | EntrezID | Gain | Loss | Normal | Detail |
|---|---|---|---|---|---|---|
| GSE15526 | CHEK2 | 11200 | 3 | 3 | 24 | |
| GSE20123 | CHEK2 | 11200 | 3 | 2 | 25 | |
| GSE43470 | CHEK2 | 11200 | 3 | 6 | 34 | |
| GSE46452 | CHEK2 | 11200 | 31 | 1 | 27 | |
| GSE47630 | CHEK2 | 11200 | 8 | 5 | 27 | |
| GSE54993 | CHEK2 | 11200 | 3 | 6 | 61 | |
| GSE54994 | CHEK2 | 11200 | 11 | 8 | 34 | |
| GSE60625 | CHEK2 | 11200 | 5 | 0 | 6 | |
| GSE74703 | CHEK2 | 11200 | 3 | 5 | 28 | |
| GSE74704 | CHEK2 | 11200 | 0 | 1 | 19 | |
| TCGA | CHEK2 | 11200 | 27 | 15 | 54 |
Total number of gains: 97; Total number of losses: 52; Total Number of normals: 339.
Somatic mutations of CHEK2:
Generating mutation plots.
Highly correlated genes for CHEK2:
Showing top 20/845 corelated genes with mean PCC>0.5.
| Gene1 | Gene2 | Mean PCC | Num. Datasets | Num. PCC<0 | Num. PCC>0.5 |
|---|---|---|---|---|---|
| CHEK2 | MCM2 | 0.748734 | 11 | 0 | 10 |
| CHEK2 | UBE2C | 0.741867 | 10 | 0 | 10 |
| CHEK2 | DNMT1 | 0.72263 | 10 | 0 | 9 |
| CHEK2 | CDCA8 | 0.72034 | 11 | 0 | 10 |
| CHEK2 | LMNB2 | 0.717358 | 10 | 0 | 9 |
| CHEK2 | IPO9 | 0.716464 | 7 | 0 | 7 |
| CHEK2 | LAMTOR4 | 0.710944 | 5 | 0 | 5 |
| CHEK2 | LRRC45 | 0.710196 | 4 | 0 | 4 |
| CHEK2 | BCR | 0.706298 | 3 | 0 | 3 |
| CHEK2 | CDCA2 | 0.706102 | 4 | 0 | 4 |
| CHEK2 | HJURP | 0.704788 | 11 | 0 | 10 |
| CHEK2 | NUSAP1 | 0.702877 | 10 | 0 | 9 |
| CHEK2 | CEP55 | 0.702607 | 11 | 0 | 10 |
| CHEK2 | KIF4A | 0.70134 | 12 | 0 | 11 |
| CHEK2 | UMPS | 0.700368 | 8 | 0 | 7 |
| CHEK2 | FOXM1 | 0.699562 | 12 | 0 | 10 |
| CHEK2 | CDC20 | 0.699266 | 11 | 0 | 10 |
| CHEK2 | PRR15 | 0.696251 | 4 | 0 | 4 |
| CHEK2 | DLGAP5 | 0.69266 | 11 | 0 | 10 |
| CHEK2 | KIF18B | 0.692488 | 10 | 0 | 9 |
For details and further investigation, click here