| Full name: cathepsin L | Alias Symbol: FLJ31037 | ||
| Type: protein-coding gene | Cytoband: 9q21.33 | ||
| Entrez ID: 1514 | HGNC ID: HGNC:2537 | Ensembl Gene: ENSG00000135047 | OMIM ID: 116880 |
| Related drugs: CHEMBL222649... [more] | |||
CTSL involved pathways:
| KEGG pathway | Description | View |
|---|---|---|
| hsa04210 | Apoptosis | |
| hsa04612 | Antigen processing and presentation |
Expression of CTSL:
| Dataset | Gene | EntrezID | Probe | Log2FC | Adj.pValue | Expression |
|---|---|---|---|---|---|---|
| GSE17351 | CTSL | 1514 | 202087_s_at | 0.5101 | 0.4707 | |
| GSE20347 | CTSL | 1514 | 202087_s_at | 0.9854 | 0.0001 | |
| GSE23400 | CTSL | 1514 | 202087_s_at | 0.7192 | 0.0000 | |
| GSE26886 | CTSL | 1514 | 202087_s_at | 1.1667 | 0.0002 | |
| GSE29001 | CTSL | 1514 | 202087_s_at | 1.8829 | 0.0000 | |
| GSE38129 | CTSL | 1514 | 202087_s_at | 0.7434 | 0.0060 | |
| GSE45670 | CTSL | 1514 | 202087_s_at | 0.3235 | 0.4484 | |
| GSE53622 | CTSL | 1514 | 35599 | 0.4114 | 0.0000 | |
| GSE53624 | CTSL | 1514 | 35599 | 0.4696 | 0.0000 | |
| GSE63941 | CTSL | 1514 | 202087_s_at | -2.4548 | 0.0056 | |
| GSE77861 | CTSL | 1514 | 202087_s_at | 1.1954 | 0.0042 | |
| SRP007169 | CTSL | 1514 | RNAseq | 4.1949 | 0.0000 | |
| SRP008496 | CTSL | 1514 | RNAseq | 4.2915 | 0.0000 | |
| SRP064894 | CTSL | 1514 | RNAseq | 1.0418 | 0.0028 | |
| SRP133303 | CTSL | 1514 | RNAseq | 1.7433 | 0.0000 | |
| SRP159526 | CTSL | 1514 | RNAseq | 1.4609 | 0.0030 | |
| SRP193095 | CTSL | 1514 | RNAseq | 3.1550 | 0.0000 | |
| SRP219564 | CTSL | 1514 | RNAseq | 0.9214 | 0.1730 |
Upregulated datasets: 9; Downregulated datasets: 1.
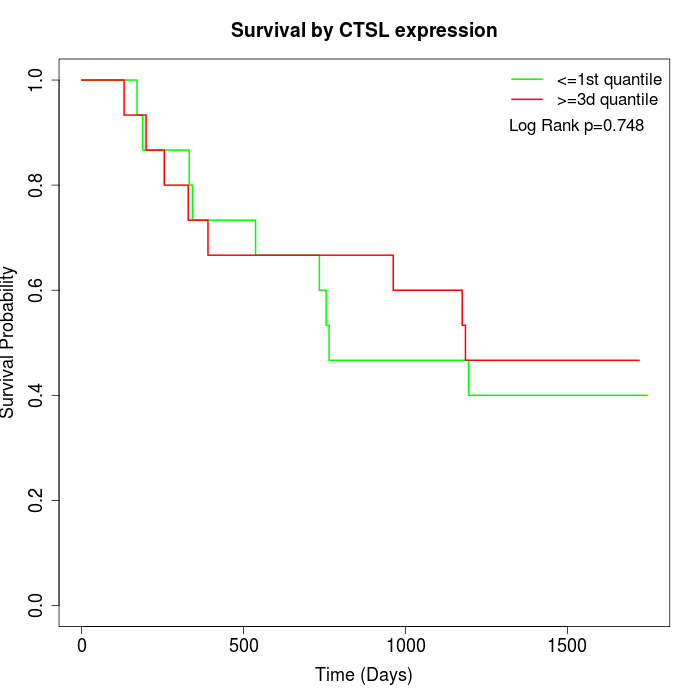
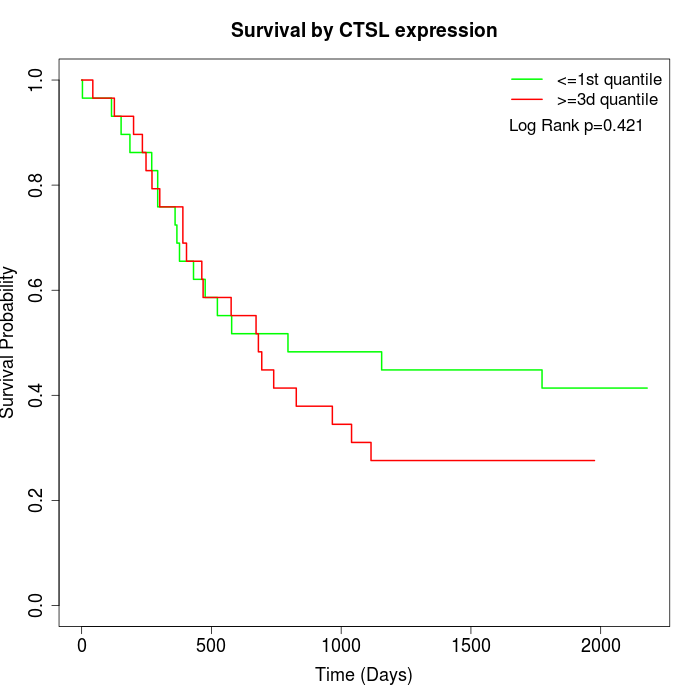
Survival by CTSL expression:
Note: Click image to view full size file.
Copy number change of CTSL:
| Dataset | Gene | EntrezID | Gain | Loss | Normal | Detail |
|---|---|---|---|---|---|---|
| GSE15526 | CTSL | 1514 | 5 | 10 | 15 | |
| GSE20123 | CTSL | 1514 | 5 | 10 | 15 | |
| GSE43470 | CTSL | 1514 | 6 | 3 | 34 | |
| GSE46452 | CTSL | 1514 | 6 | 14 | 39 | |
| GSE47630 | CTSL | 1514 | 1 | 18 | 21 | |
| GSE54993 | CTSL | 1514 | 4 | 2 | 64 | |
| GSE54994 | CTSL | 1514 | 6 | 11 | 36 | |
| GSE60625 | CTSL | 1514 | 0 | 0 | 11 | |
| GSE74703 | CTSL | 1514 | 5 | 3 | 28 | |
| GSE74704 | CTSL | 1514 | 3 | 7 | 10 | |
| TCGA | CTSL | 1514 | 20 | 26 | 50 |
Total number of gains: 61; Total number of losses: 104; Total Number of normals: 323.
Somatic mutations of CTSL:
Generating mutation plots.
Highly correlated genes for CTSL:
Showing top 20/907 corelated genes with mean PCC>0.5.
| Gene1 | Gene2 | Mean PCC | Num. Datasets | Num. PCC<0 | Num. PCC>0.5 |
|---|---|---|---|---|---|
| CTSL | LGALS1 | 0.742399 | 10 | 0 | 10 |
| CTSL | ASH1L-AS1 | 0.741272 | 3 | 0 | 3 |
| CTSL | LHFPL2 | 0.73262 | 11 | 0 | 11 |
| CTSL | FAP | 0.723959 | 9 | 0 | 9 |
| CTSL | COL5A1 | 0.707615 | 11 | 0 | 11 |
| CTSL | EML1 | 0.703767 | 3 | 0 | 3 |
| CTSL | RPP25 | 0.698357 | 3 | 0 | 3 |
| CTSL | IFITM1 | 0.697637 | 8 | 0 | 8 |
| CTSL | CSGALNACT2 | 0.697387 | 11 | 0 | 10 |
| CTSL | HEXA | 0.697105 | 7 | 0 | 7 |
| CTSL | INHBA | 0.696605 | 11 | 0 | 10 |
| CTSL | COL6A3 | 0.696556 | 11 | 0 | 11 |
| CTSL | VCAN | 0.694984 | 11 | 0 | 10 |
| CTSL | MMD | 0.688606 | 11 | 0 | 10 |
| CTSL | LPCAT1 | 0.686313 | 11 | 0 | 10 |
| CTSL | CHN1 | 0.685766 | 11 | 0 | 10 |
| CTSL | HEXB | 0.685533 | 11 | 0 | 11 |
| CTSL | NID2 | 0.683882 | 9 | 0 | 8 |
| CTSL | COL4A1 | 0.683863 | 11 | 0 | 10 |
| CTSL | ARPC1B | 0.681509 | 10 | 0 | 8 |
For details and further investigation, click here