| Full name: four and a half LIM domains 2 | Alias Symbol: SLIM3|DRAL | ||
| Type: protein-coding gene | Cytoband: 2q12.2 | ||
| Entrez ID: 2274 | HGNC ID: HGNC:3703 | Ensembl Gene: ENSG00000115641 | OMIM ID: 602633 |
FHL2 involved pathways:
| KEGG pathway | Description | View |
|---|---|---|
| hsa04380 | Osteoclast differentiation |
Expression of FHL2:
| Dataset | Gene | EntrezID | Probe | Log2FC | Adj.pValue | Expression |
|---|---|---|---|---|---|---|
| GSE17351 | FHL2 | 2274 | 202949_s_at | 0.0220 | 0.9826 | |
| GSE20347 | FHL2 | 2274 | 202949_s_at | 0.4636 | 0.1280 | |
| GSE23400 | FHL2 | 2274 | 202949_s_at | 0.4808 | 0.0001 | |
| GSE26886 | FHL2 | 2274 | 202949_s_at | 1.1153 | 0.0040 | |
| GSE29001 | FHL2 | 2274 | 202949_s_at | 0.1739 | 0.7327 | |
| GSE38129 | FHL2 | 2274 | 202949_s_at | 0.3878 | 0.0848 | |
| GSE45670 | FHL2 | 2274 | 202949_s_at | -0.2191 | 0.6223 | |
| GSE53622 | FHL2 | 2274 | 98457 | 0.1789 | 0.0714 | |
| GSE53624 | FHL2 | 2274 | 98457 | 0.2951 | 0.0016 | |
| GSE63941 | FHL2 | 2274 | 202949_s_at | -2.6265 | 0.0132 | |
| GSE77861 | FHL2 | 2274 | 202949_s_at | 0.3337 | 0.5253 | |
| GSE97050 | FHL2 | 2274 | A_23_P108751 | 0.2729 | 0.2594 | |
| SRP007169 | FHL2 | 2274 | RNAseq | 1.8469 | 0.0005 | |
| SRP008496 | FHL2 | 2274 | RNAseq | 2.4172 | 0.0000 | |
| SRP064894 | FHL2 | 2274 | RNAseq | 0.3187 | 0.1870 | |
| SRP133303 | FHL2 | 2274 | RNAseq | 0.7261 | 0.0141 | |
| SRP159526 | FHL2 | 2274 | RNAseq | -0.0213 | 0.9581 | |
| SRP193095 | FHL2 | 2274 | RNAseq | -0.1746 | 0.5332 | |
| SRP219564 | FHL2 | 2274 | RNAseq | -0.1109 | 0.8123 | |
| TCGA | FHL2 | 2274 | RNAseq | 0.0044 | 0.9593 |
Upregulated datasets: 3; Downregulated datasets: 1.
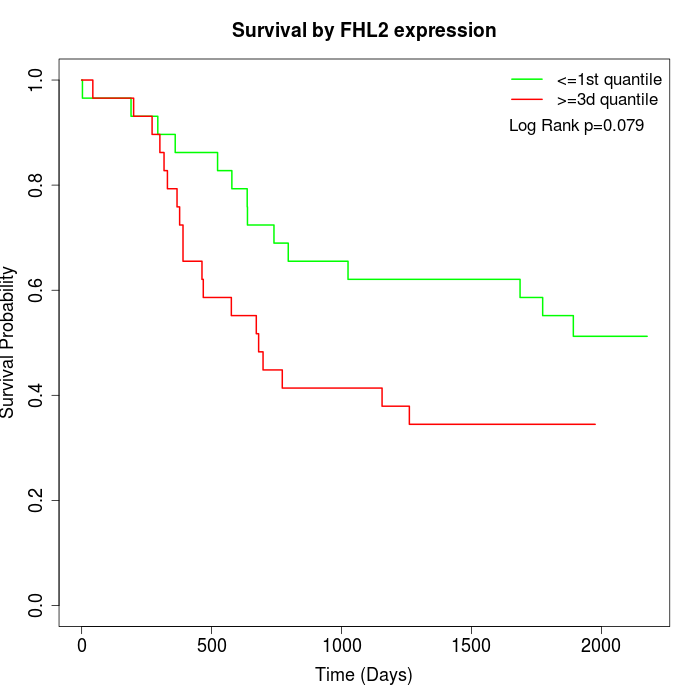
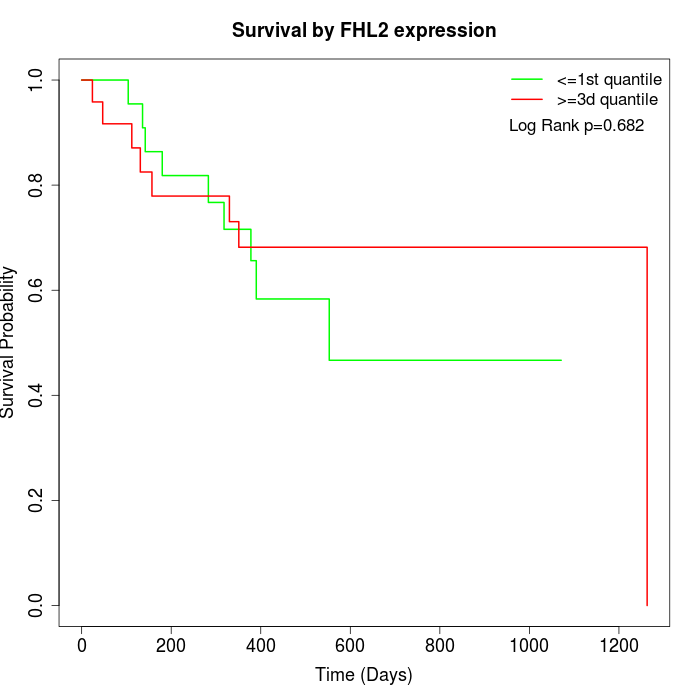
Survival by FHL2 expression:
Note: Click image to view full size file.
Copy number change of FHL2:
| Dataset | Gene | EntrezID | Gain | Loss | Normal | Detail |
|---|---|---|---|---|---|---|
| GSE15526 | FHL2 | 2274 | 3 | 2 | 25 | |
| GSE20123 | FHL2 | 2274 | 3 | 2 | 25 | |
| GSE43470 | FHL2 | 2274 | 3 | 1 | 39 | |
| GSE46452 | FHL2 | 2274 | 1 | 4 | 54 | |
| GSE47630 | FHL2 | 2274 | 7 | 0 | 33 | |
| GSE54993 | FHL2 | 2274 | 0 | 6 | 64 | |
| GSE54994 | FHL2 | 2274 | 10 | 0 | 43 | |
| GSE60625 | FHL2 | 2274 | 0 | 3 | 8 | |
| GSE74703 | FHL2 | 2274 | 3 | 1 | 32 | |
| GSE74704 | FHL2 | 2274 | 2 | 2 | 16 | |
| TCGA | FHL2 | 2274 | 35 | 5 | 56 |
Total number of gains: 67; Total number of losses: 26; Total Number of normals: 395.
Somatic mutations of FHL2:
Generating mutation plots.
Highly correlated genes for FHL2:
Showing top 20/603 corelated genes with mean PCC>0.5.
| Gene1 | Gene2 | Mean PCC | Num. Datasets | Num. PCC<0 | Num. PCC>0.5 |
|---|---|---|---|---|---|
| FHL2 | MTX3 | 0.786749 | 3 | 0 | 3 |
| FHL2 | PHRF1 | 0.725136 | 3 | 0 | 3 |
| FHL2 | IRF7 | 0.708335 | 3 | 0 | 3 |
| FHL2 | FAM160B1 | 0.699762 | 3 | 0 | 3 |
| FHL2 | PTPN23 | 0.698414 | 3 | 0 | 3 |
| FHL2 | MRC2 | 0.698273 | 3 | 0 | 3 |
| FHL2 | RGL1 | 0.695973 | 3 | 0 | 3 |
| FHL2 | HEATR6 | 0.692348 | 3 | 0 | 3 |
| FHL2 | TINF2 | 0.68738 | 4 | 0 | 4 |
| FHL2 | RING1 | 0.687094 | 3 | 0 | 3 |
| FHL2 | CARD10 | 0.67237 | 7 | 0 | 6 |
| FHL2 | PLEKHA4 | 0.67092 | 4 | 0 | 4 |
| FHL2 | RSPO3 | 0.66356 | 3 | 0 | 3 |
| FHL2 | MRGPRF | 0.66178 | 3 | 0 | 3 |
| FHL2 | ASPH | 0.657043 | 3 | 0 | 3 |
| FHL2 | VGLL3 | 0.656141 | 6 | 0 | 4 |
| FHL2 | ANXA4 | 0.655267 | 7 | 0 | 5 |
| FHL2 | GLT8D1 | 0.653864 | 5 | 0 | 4 |
| FHL2 | NLE1 | 0.651543 | 3 | 0 | 3 |
| FHL2 | MAP4 | 0.648833 | 4 | 0 | 3 |
For details and further investigation, click here