| Full name: glucosaminyl (N-acetyl) transferase family member 7 | Alias Symbol: dJ1153D9.2 | ||
| Type: protein-coding gene | Cytoband: 20q13.31 | ||
| Entrez ID: 140687 | HGNC ID: HGNC:16099 | Ensembl Gene: ENSG00000124091 | OMIM ID: |
Expression of GCNT7:
| Dataset | Gene | EntrezID | Probe | Log2FC | Adj.pValue | Expression |
|---|---|---|---|---|---|---|
| GSE17351 | GCNT7 | 140687 | 242040_at | 0.5010 | 0.2996 | |
| GSE26886 | GCNT7 | 140687 | 242040_at | 0.1992 | 0.2026 | |
| GSE45670 | GCNT7 | 140687 | 242040_at | 0.3292 | 0.0229 | |
| GSE53622 | GCNT7 | 140687 | 104942 | 0.2326 | 0.0963 | |
| GSE53624 | GCNT7 | 140687 | 104942 | 0.4791 | 0.0000 | |
| GSE63941 | GCNT7 | 140687 | 242040_at | -0.1754 | 0.4363 | |
| GSE77861 | GCNT7 | 140687 | 242040_at | -0.0769 | 0.5910 | |
| GSE97050 | GCNT7 | 140687 | A_33_P3282522 | 0.1144 | 0.5695 | |
| SRP133303 | GCNT7 | 140687 | RNAseq | 0.3313 | 0.0957 | |
| TCGA | GCNT7 | 140687 | RNAseq | -0.1015 | 0.7913 |
Upregulated datasets: 0; Downregulated datasets: 0.
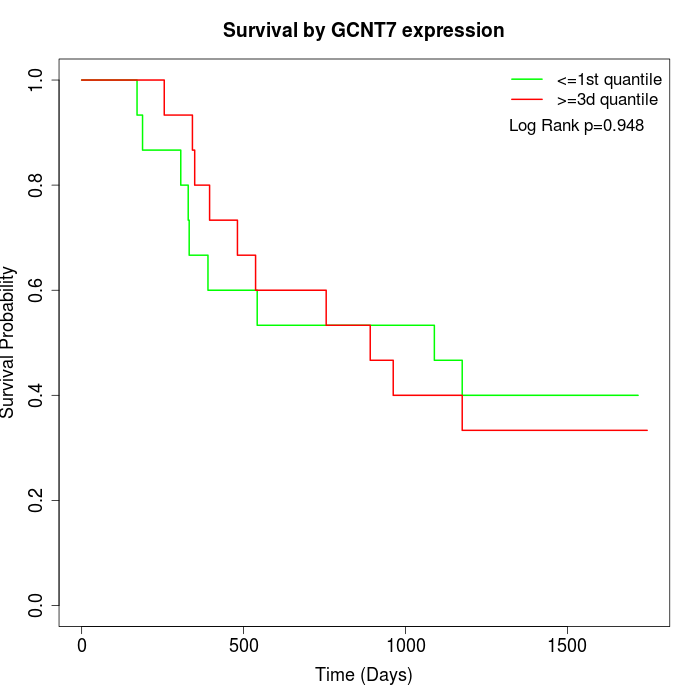
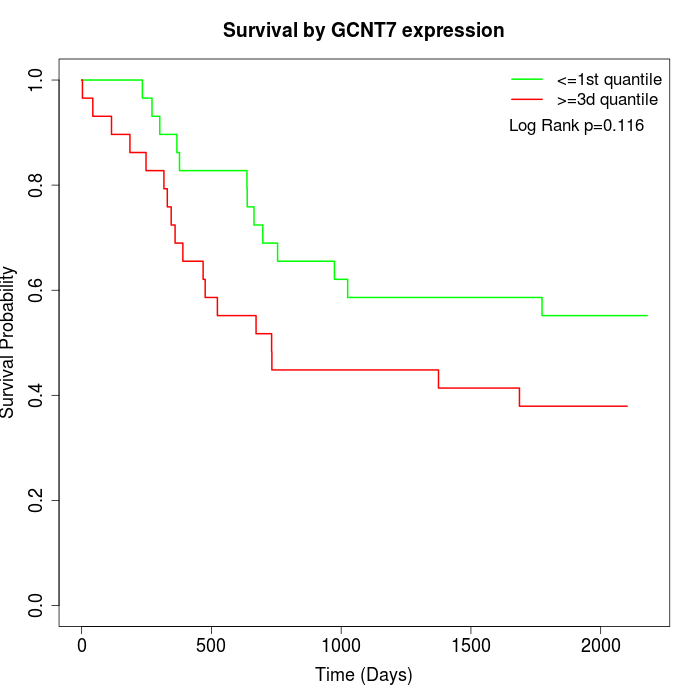
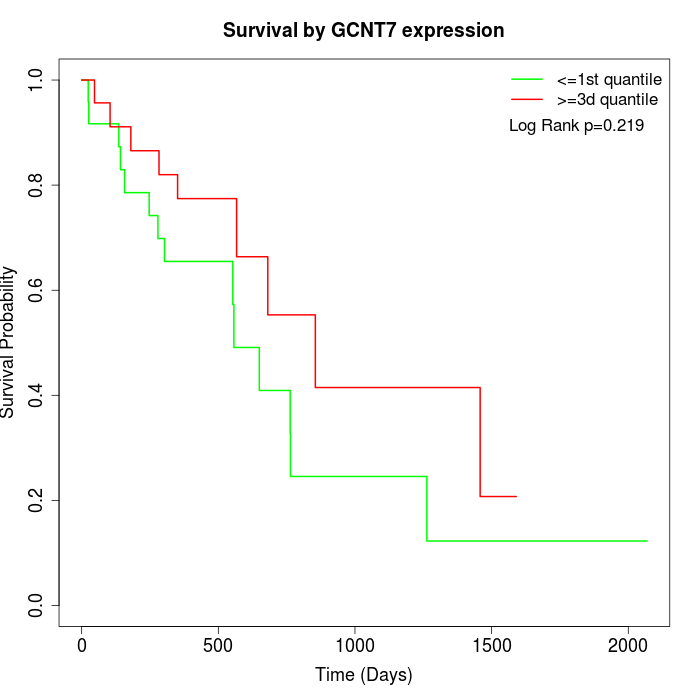
Survival by GCNT7 expression:
Note: Click image to view full size file.
Copy number change of GCNT7:
| Dataset | Gene | EntrezID | Gain | Loss | Normal | Detail |
|---|---|---|---|---|---|---|
| GSE15526 | GCNT7 | 140687 | 14 | 2 | 14 | |
| GSE20123 | GCNT7 | 140687 | 14 | 2 | 14 | |
| GSE43470 | GCNT7 | 140687 | 13 | 0 | 30 | |
| GSE46452 | GCNT7 | 140687 | 29 | 0 | 30 | |
| GSE47630 | GCNT7 | 140687 | 24 | 1 | 15 | |
| GSE54993 | GCNT7 | 140687 | 0 | 17 | 53 | |
| GSE54994 | GCNT7 | 140687 | 27 | 0 | 26 | |
| GSE60625 | GCNT7 | 140687 | 0 | 0 | 11 | |
| GSE74703 | GCNT7 | 140687 | 11 | 0 | 25 | |
| GSE74704 | GCNT7 | 140687 | 11 | 0 | 9 | |
| TCGA | GCNT7 | 140687 | 43 | 4 | 49 |
Total number of gains: 186; Total number of losses: 26; Total Number of normals: 276.
Somatic mutations of GCNT7:
Generating mutation plots.
Highly correlated genes for GCNT7:
Showing top 20/23 corelated genes with mean PCC>0.5.
| Gene1 | Gene2 | Mean PCC | Num. Datasets | Num. PCC<0 | Num. PCC>0.5 |
|---|---|---|---|---|---|
| GCNT7 | LRRC8E | 0.687147 | 3 | 0 | 3 |
| GCNT7 | BATF2 | 0.685823 | 3 | 0 | 3 |
| GCNT7 | ZNF556 | 0.652802 | 3 | 0 | 3 |
| GCNT7 | LY6K | 0.620261 | 3 | 0 | 3 |
| GCNT7 | SIPA1L3 | 0.620118 | 3 | 0 | 3 |
| GCNT7 | ADRM1 | 0.609906 | 3 | 0 | 3 |
| GCNT7 | MANF | 0.598357 | 3 | 0 | 3 |
| GCNT7 | OGFR | 0.596522 | 4 | 0 | 3 |
| GCNT7 | B4GALT7 | 0.591544 | 4 | 0 | 3 |
| GCNT7 | FAM91A1 | 0.590976 | 3 | 0 | 3 |
| GCNT7 | SH3D21 | 0.581383 | 3 | 0 | 3 |
| GCNT7 | LPA | 0.578477 | 4 | 0 | 3 |
| GCNT7 | KMO | 0.57687 | 3 | 0 | 3 |
| GCNT7 | SIRT6 | 0.572939 | 3 | 0 | 3 |
| GCNT7 | MALAT1 | 0.565159 | 3 | 0 | 3 |
| GCNT7 | MICALL2 | 0.562493 | 4 | 0 | 3 |
| GCNT7 | CPNE1 | 0.544418 | 3 | 0 | 3 |
| GCNT7 | KCNQ1OT1 | 0.544211 | 5 | 0 | 3 |
| GCNT7 | TTI1 | 0.540909 | 4 | 0 | 3 |
| GCNT7 | CCDC59 | 0.537871 | 3 | 0 | 3 |
For details and further investigation, click here