| Full name: mitogen-activated protein kinase kinase 4 | Alias Symbol: MEK4|JNKK1|PRKMK4|MKK4 | ||
| Type: protein-coding gene | Cytoband: 17p12 | ||
| Entrez ID: 6416 | HGNC ID: HGNC:6844 | Ensembl Gene: ENSG00000065559 | OMIM ID: 601335 |
| Related drugs: BINIMETINIB, COBIMETINIB, DABRAFENIB MESYLATE, LEUCOVORIN, PD-0325901, Refametinib, SELUMETINIB, STAUROSPORINE, TRAMETINIB, WX-554... [more] | |||
Screen Evidence:
| |||
MAP2K4 involved pathways:
Expression of MAP2K4:
| Dataset | Gene | EntrezID | Probe | Log2FC | Adj.pValue | Expression |
|---|---|---|---|---|---|---|
| GSE17351 | MAP2K4 | 6416 | 203266_s_at | -0.3556 | 0.5465 | |
| GSE20347 | MAP2K4 | 6416 | 203266_s_at | -0.7415 | 0.0001 | |
| GSE23400 | MAP2K4 | 6416 | 203266_s_at | -0.3788 | 0.0000 | |
| GSE26886 | MAP2K4 | 6416 | 203266_s_at | -0.8212 | 0.0001 | |
| GSE29001 | MAP2K4 | 6416 | 203266_s_at | -0.5658 | 0.0195 | |
| GSE38129 | MAP2K4 | 6416 | 203266_s_at | -0.4373 | 0.0077 | |
| GSE45670 | MAP2K4 | 6416 | 203266_s_at | -0.1376 | 0.2752 | |
| GSE53622 | MAP2K4 | 6416 | 3319 | -0.7689 | 0.0000 | |
| GSE53624 | MAP2K4 | 6416 | 3319 | -0.6577 | 0.0000 | |
| GSE63941 | MAP2K4 | 6416 | 203266_s_at | -0.2963 | 0.5331 | |
| GSE77861 | MAP2K4 | 6416 | 203266_s_at | -0.2212 | 0.2612 | |
| GSE97050 | MAP2K4 | 6416 | A_33_P3653330 | 0.1230 | 0.6291 | |
| SRP007169 | MAP2K4 | 6416 | RNAseq | -1.3031 | 0.0061 | |
| SRP008496 | MAP2K4 | 6416 | RNAseq | -1.0064 | 0.0000 | |
| SRP064894 | MAP2K4 | 6416 | RNAseq | -0.4034 | 0.1333 | |
| SRP133303 | MAP2K4 | 6416 | RNAseq | -0.4102 | 0.0045 | |
| SRP159526 | MAP2K4 | 6416 | RNAseq | -0.8057 | 0.0078 | |
| SRP193095 | MAP2K4 | 6416 | RNAseq | -0.7395 | 0.0000 | |
| SRP219564 | MAP2K4 | 6416 | RNAseq | -0.6174 | 0.0664 | |
| TCGA | MAP2K4 | 6416 | RNAseq | -0.0558 | 0.2464 |
Upregulated datasets: 0; Downregulated datasets: 2.
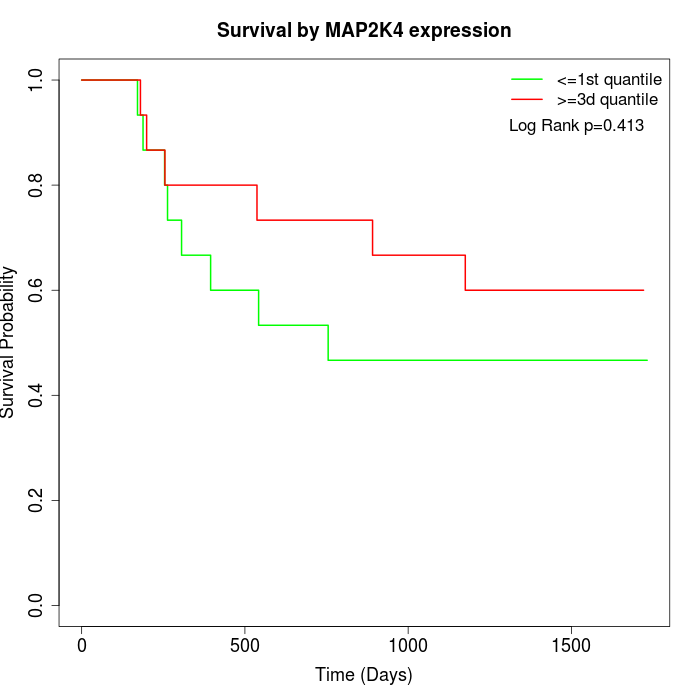
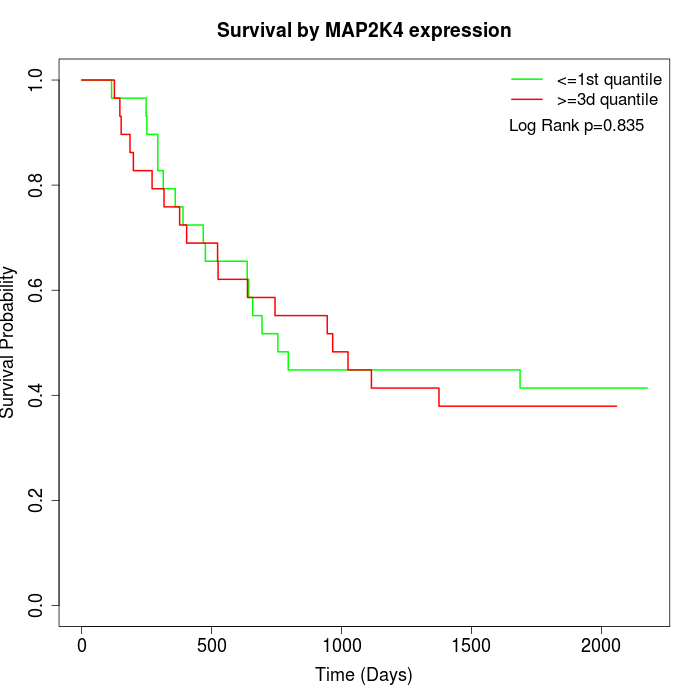
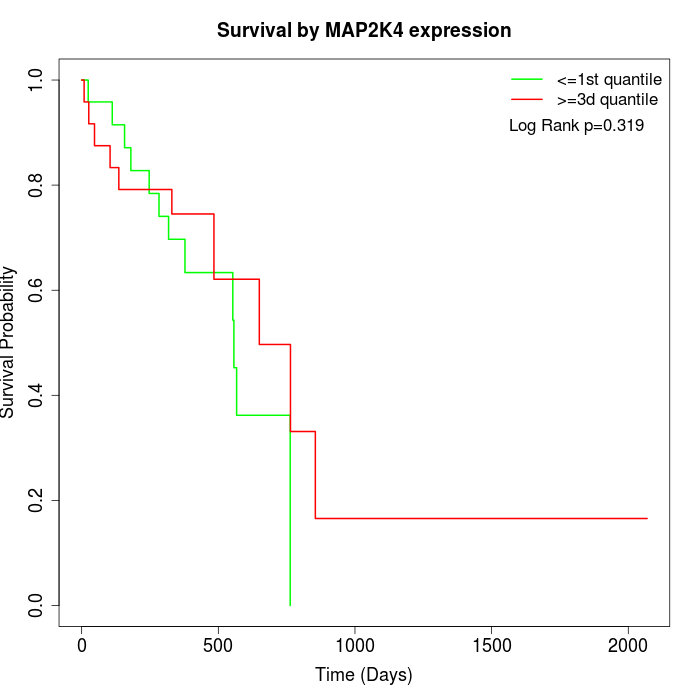
Survival by MAP2K4 expression:
Note: Click image to view full size file.
Copy number change of MAP2K4:
| Dataset | Gene | EntrezID | Gain | Loss | Normal | Detail |
|---|---|---|---|---|---|---|
| GSE15526 | MAP2K4 | 6416 | 3 | 4 | 23 | |
| GSE20123 | MAP2K4 | 6416 | 3 | 5 | 22 | |
| GSE43470 | MAP2K4 | 6416 | 2 | 5 | 36 | |
| GSE46452 | MAP2K4 | 6416 | 34 | 1 | 24 | |
| GSE47630 | MAP2K4 | 6416 | 7 | 1 | 32 | |
| GSE54993 | MAP2K4 | 6416 | 3 | 2 | 65 | |
| GSE54994 | MAP2K4 | 6416 | 6 | 6 | 41 | |
| GSE60625 | MAP2K4 | 6416 | 4 | 0 | 7 | |
| GSE74703 | MAP2K4 | 6416 | 2 | 4 | 30 | |
| GSE74704 | MAP2K4 | 6416 | 2 | 2 | 16 | |
| TCGA | MAP2K4 | 6416 | 18 | 21 | 57 |
Total number of gains: 84; Total number of losses: 51; Total Number of normals: 353.
Somatic mutations of MAP2K4:
Generating mutation plots.
Highly correlated genes for MAP2K4:
Showing top 20/1206 corelated genes with mean PCC>0.5.
| Gene1 | Gene2 | Mean PCC | Num. Datasets | Num. PCC<0 | Num. PCC>0.5 |
|---|---|---|---|---|---|
| MAP2K4 | RNFT1 | 0.7611 | 3 | 0 | 3 |
| MAP2K4 | NUAK2 | 0.754344 | 3 | 0 | 3 |
| MAP2K4 | NIPAL1 | 0.732154 | 3 | 0 | 3 |
| MAP2K4 | TAX1BP3 | 0.711437 | 3 | 0 | 3 |
| MAP2K4 | ZNF823 | 0.708579 | 5 | 0 | 5 |
| MAP2K4 | MPZL3 | 0.698255 | 7 | 0 | 6 |
| MAP2K4 | ACOT12 | 0.697419 | 3 | 0 | 3 |
| MAP2K4 | KIF21A | 0.688968 | 5 | 0 | 4 |
| MAP2K4 | LTA4H | 0.687362 | 3 | 0 | 3 |
| MAP2K4 | SH3GL1 | 0.685739 | 11 | 0 | 10 |
| MAP2K4 | TBC1D2 | 0.683817 | 4 | 0 | 3 |
| MAP2K4 | GBP6 | 0.681998 | 4 | 0 | 4 |
| MAP2K4 | ZDHHC12 | 0.681263 | 3 | 0 | 3 |
| MAP2K4 | SIAE | 0.681038 | 5 | 0 | 4 |
| MAP2K4 | CAPN14 | 0.679949 | 5 | 0 | 4 |
| MAP2K4 | ANKRD31 | 0.675692 | 4 | 0 | 4 |
| MAP2K4 | C7orf57 | 0.671374 | 5 | 0 | 5 |
| MAP2K4 | PLIN3 | 0.671016 | 10 | 0 | 10 |
| MAP2K4 | FRMD4B | 0.669664 | 10 | 0 | 9 |
| MAP2K4 | GSTZ1 | 0.669365 | 4 | 0 | 3 |
For details and further investigation, click here