| Full name: nuclear receptor coactivator 4 | Alias Symbol: ARA70|RFG|ELE1|PTC3|DKFZp762E1112 | ||
| Type: protein-coding gene | Cytoband: 10q11.22 | ||
| Entrez ID: 8031 | HGNC ID: HGNC:7671 | Ensembl Gene: ENSG00000266412 | OMIM ID: 601984 |
Screen Evidence:
| |||
NCOA4 involved pathways:
| KEGG pathway | Description | View |
|---|---|---|
| hsa05200 | Pathways in cancer | |
| hsa05216 | Thyroid cancer |
Expression of NCOA4:
| Dataset | Gene | EntrezID | Probe | Log2FC | Adj.pValue | Expression |
|---|---|---|---|---|---|---|
| GSE17351 | NCOA4 | 8031 | 210774_s_at | -0.4286 | 0.2333 | |
| GSE20347 | NCOA4 | 8031 | 210774_s_at | -0.5921 | 0.0000 | |
| GSE23400 | NCOA4 | 8031 | 210774_s_at | -0.3404 | 0.0000 | |
| GSE26886 | NCOA4 | 8031 | 210774_s_at | -1.0066 | 0.0000 | |
| GSE29001 | NCOA4 | 8031 | 210774_s_at | -0.4281 | 0.0130 | |
| GSE38129 | NCOA4 | 8031 | 210774_s_at | -0.5461 | 0.0000 | |
| GSE45670 | NCOA4 | 8031 | 210774_s_at | -0.3116 | 0.0020 | |
| GSE53622 | NCOA4 | 8031 | 3413 | -0.8150 | 0.0000 | |
| GSE53624 | NCOA4 | 8031 | 3413 | -0.4066 | 0.0000 | |
| GSE63941 | NCOA4 | 8031 | 210774_s_at | -0.4771 | 0.1916 | |
| GSE77861 | NCOA4 | 8031 | 210774_s_at | -0.5391 | 0.2630 | |
| GSE97050 | NCOA4 | 8031 | A_33_P3270034 | -0.3461 | 0.2017 | |
| SRP007169 | NCOA4 | 8031 | RNAseq | -0.2614 | 0.5656 | |
| SRP008496 | NCOA4 | 8031 | RNAseq | -0.1751 | 0.5488 | |
| SRP064894 | NCOA4 | 8031 | RNAseq | -0.8767 | 0.0000 | |
| SRP133303 | NCOA4 | 8031 | RNAseq | -0.2958 | 0.0294 | |
| SRP159526 | NCOA4 | 8031 | RNAseq | -0.9579 | 0.0561 | |
| SRP193095 | NCOA4 | 8031 | RNAseq | -0.8589 | 0.0000 | |
| SRP219564 | NCOA4 | 8031 | RNAseq | -0.3701 | 0.2349 | |
| TCGA | NCOA4 | 8031 | RNAseq | -0.0973 | 0.0141 |
Upregulated datasets: 0; Downregulated datasets: 1.
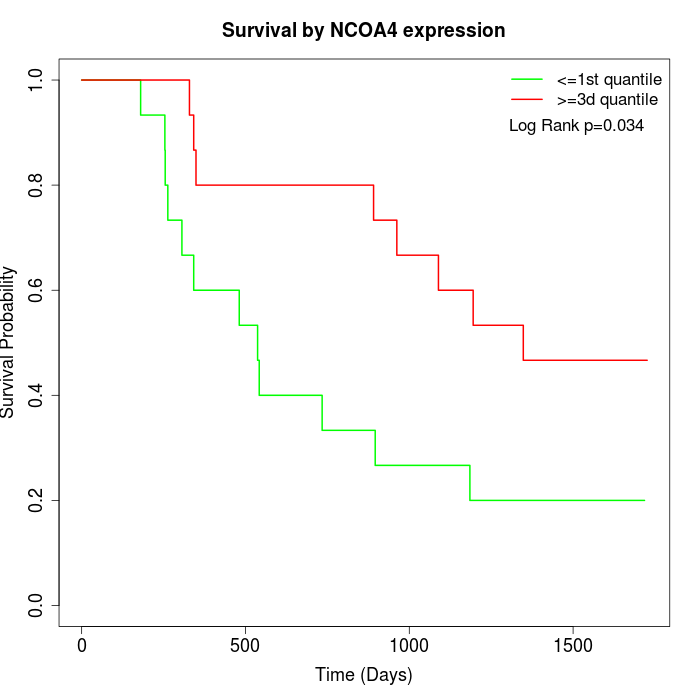
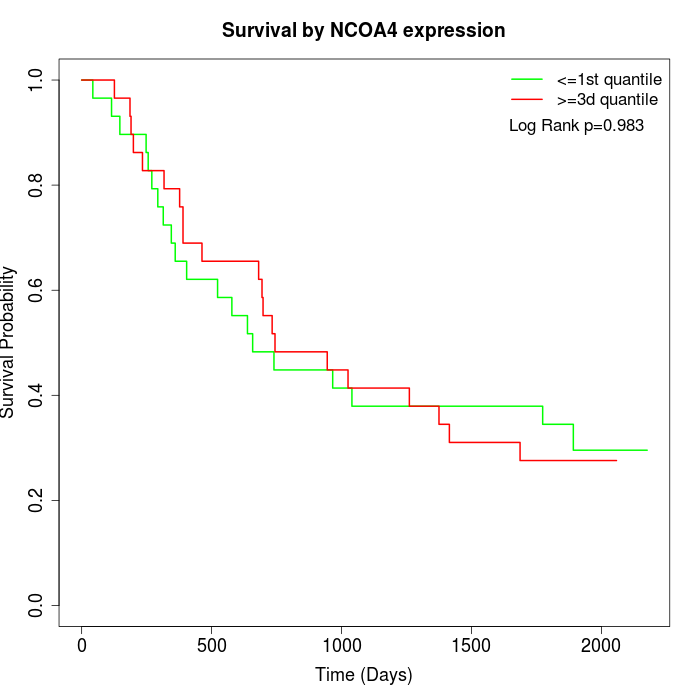
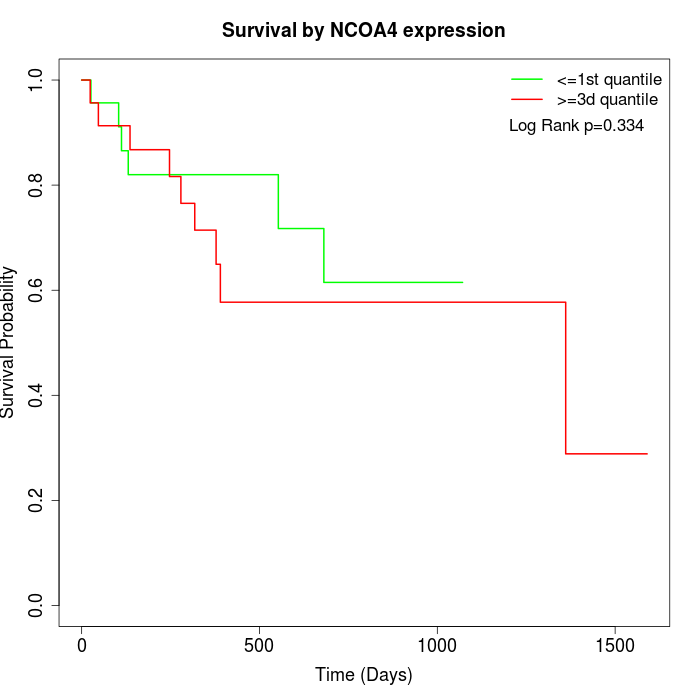
Survival by NCOA4 expression:
Note: Click image to view full size file.
Copy number change of NCOA4:
| Dataset | Gene | EntrezID | Gain | Loss | Normal | Detail |
|---|---|---|---|---|---|---|
| GSE15526 | NCOA4 | 8031 | 2 | 5 | 23 | |
| GSE20123 | NCOA4 | 8031 | 2 | 5 | 23 | |
| GSE43470 | NCOA4 | 8031 | 2 | 6 | 35 | |
| GSE46452 | NCOA4 | 8031 | 0 | 13 | 46 | |
| GSE47630 | NCOA4 | 8031 | 4 | 12 | 24 | |
| GSE54993 | NCOA4 | 8031 | 9 | 0 | 61 | |
| GSE54994 | NCOA4 | 8031 | 2 | 10 | 41 | |
| GSE60625 | NCOA4 | 8031 | 0 | 0 | 11 | |
| GSE74703 | NCOA4 | 8031 | 2 | 3 | 31 | |
| GSE74704 | NCOA4 | 8031 | 1 | 4 | 15 | |
| TCGA | NCOA4 | 8031 | 13 | 22 | 61 |
Total number of gains: 37; Total number of losses: 80; Total Number of normals: 371.
Somatic mutations of NCOA4:
Generating mutation plots.
Highly correlated genes for NCOA4:
Showing top 20/1473 corelated genes with mean PCC>0.5.
| Gene1 | Gene2 | Mean PCC | Num. Datasets | Num. PCC<0 | Num. PCC>0.5 |
|---|---|---|---|---|---|
| NCOA4 | MKRN1 | 0.77027 | 3 | 0 | 3 |
| NCOA4 | TBC1D20 | 0.770179 | 3 | 0 | 3 |
| NCOA4 | AK3 | 0.75741 | 7 | 0 | 6 |
| NCOA4 | FAM122A | 0.754526 | 4 | 0 | 4 |
| NCOA4 | SMU1 | 0.743062 | 3 | 0 | 3 |
| NCOA4 | MIER3 | 0.738903 | 3 | 0 | 3 |
| NCOA4 | IMPACT | 0.737943 | 3 | 0 | 3 |
| NCOA4 | STK38 | 0.737549 | 5 | 0 | 5 |
| NCOA4 | NDUFA4 | 0.737522 | 3 | 0 | 3 |
| NCOA4 | TRAPPC6B | 0.730907 | 3 | 0 | 3 |
| NCOA4 | CAPRIN1 | 0.730735 | 3 | 0 | 3 |
| NCOA4 | BLOC1S2 | 0.730192 | 4 | 0 | 3 |
| NCOA4 | RC3H1 | 0.719719 | 4 | 0 | 4 |
| NCOA4 | BRK1 | 0.712338 | 8 | 0 | 7 |
| NCOA4 | UQCRC2 | 0.711981 | 7 | 0 | 5 |
| NCOA4 | CETN2 | 0.711625 | 5 | 0 | 5 |
| NCOA4 | CCNY | 0.709256 | 3 | 0 | 3 |
| NCOA4 | ING4 | 0.707458 | 3 | 0 | 3 |
| NCOA4 | RPS16 | 0.707256 | 3 | 0 | 3 |
| NCOA4 | MRPS11 | 0.705034 | 3 | 0 | 3 |
For details and further investigation, click here