| Full name: p53 apoptosis effector related to PMP22 | Alias Symbol: PIGPC1|dJ496H19.1|KCP1|THW|KRTCAP1 | ||
| Type: protein-coding gene | Cytoband: 6q23.3 | ||
| Entrez ID: 64065 | HGNC ID: HGNC:17637 | Ensembl Gene: ENSG00000112378 | OMIM ID: 609301 |
Screen Evidence:
| |||
PERP involved pathways:
| KEGG pathway | Description | View |
|---|---|---|
| hsa04115 | p53 signaling pathway |
Expression of PERP:
| Dataset | Gene | EntrezID | Probe | Log2FC | Adj.pValue | Expression |
|---|---|---|---|---|---|---|
| GSE17351 | PERP | 64065 | 222392_x_at | -0.0602 | 0.8864 | |
| GSE20347 | PERP | 64065 | 217744_s_at | -0.7804 | 0.0000 | |
| GSE23400 | PERP | 64065 | 217744_s_at | -0.2055 | 0.0813 | |
| GSE26886 | PERP | 64065 | 222392_x_at | -0.5735 | 0.0005 | |
| GSE29001 | PERP | 64065 | 217744_s_at | -0.6413 | 0.0216 | |
| GSE38129 | PERP | 64065 | 217744_s_at | 0.2060 | 0.6722 | |
| GSE45670 | PERP | 64065 | 222392_x_at | 0.0848 | 0.2048 | |
| GSE53622 | PERP | 64065 | 153810 | -0.4674 | 0.0077 | |
| GSE53624 | PERP | 64065 | 153810 | -0.6559 | 0.0000 | |
| GSE63941 | PERP | 64065 | 222392_x_at | -0.2370 | 0.5994 | |
| GSE77861 | PERP | 64065 | 222392_x_at | -0.1074 | 0.5232 | |
| GSE97050 | PERP | 64065 | A_23_P214950 | 0.4915 | 0.4552 | |
| SRP007169 | PERP | 64065 | RNAseq | -2.1444 | 0.0000 | |
| SRP008496 | PERP | 64065 | RNAseq | -2.0060 | 0.0000 | |
| SRP064894 | PERP | 64065 | RNAseq | -0.9422 | 0.0099 | |
| SRP133303 | PERP | 64065 | RNAseq | -0.6550 | 0.0016 | |
| SRP159526 | PERP | 64065 | RNAseq | -0.9599 | 0.0768 | |
| SRP193095 | PERP | 64065 | RNAseq | -1.1501 | 0.0003 | |
| SRP219564 | PERP | 64065 | RNAseq | -1.5001 | 0.0560 | |
| TCGA | PERP | 64065 | RNAseq | 0.5065 | 0.0000 |
Upregulated datasets: 0; Downregulated datasets: 3.
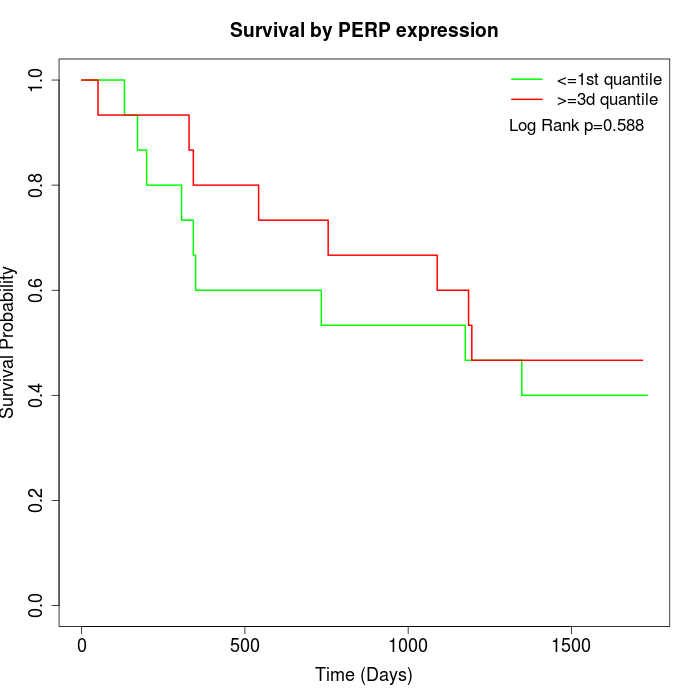
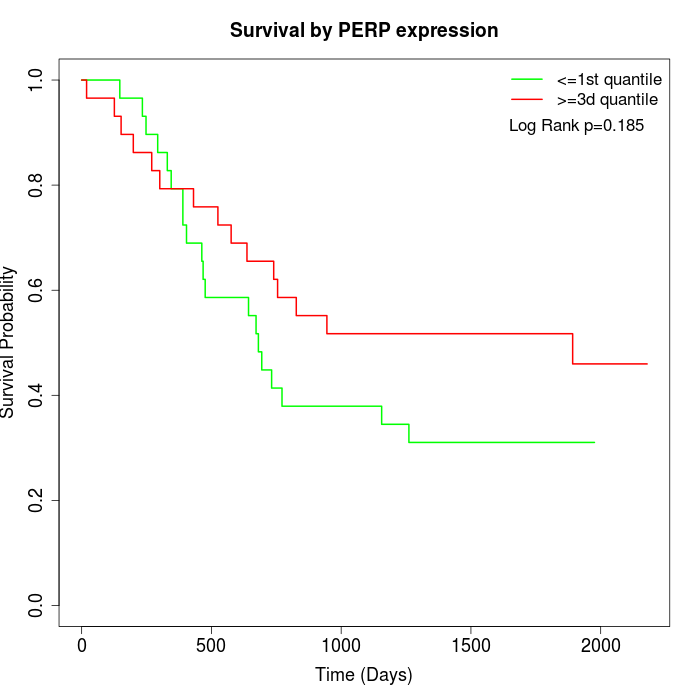
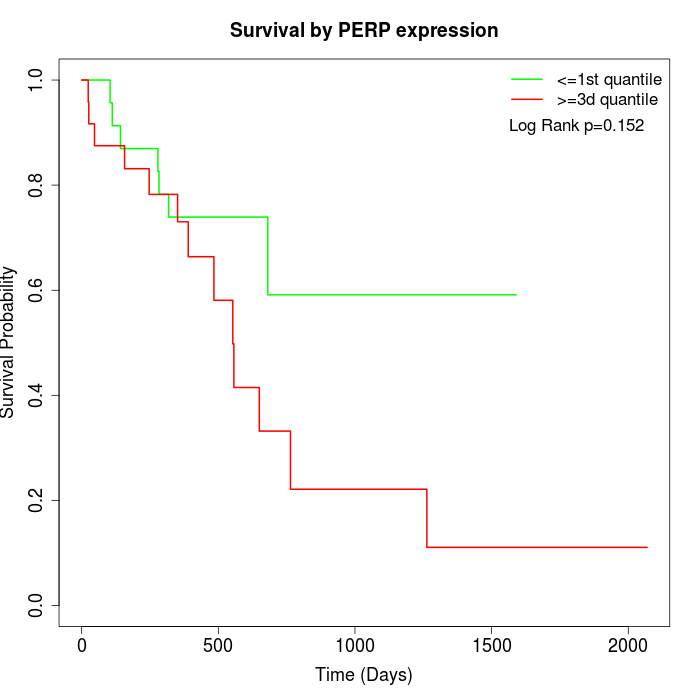
Survival by PERP expression:
Note: Click image to view full size file.
Copy number change of PERP:
| Dataset | Gene | EntrezID | Gain | Loss | Normal | Detail |
|---|---|---|---|---|---|---|
| GSE15526 | PERP | 64065 | 3 | 4 | 23 | |
| GSE20123 | PERP | 64065 | 3 | 3 | 24 | |
| GSE43470 | PERP | 64065 | 5 | 0 | 38 | |
| GSE46452 | PERP | 64065 | 3 | 10 | 46 | |
| GSE47630 | PERP | 64065 | 9 | 4 | 27 | |
| GSE54993 | PERP | 64065 | 3 | 2 | 65 | |
| GSE54994 | PERP | 64065 | 8 | 8 | 37 | |
| GSE60625 | PERP | 64065 | 0 | 1 | 10 | |
| GSE74703 | PERP | 64065 | 4 | 0 | 32 | |
| GSE74704 | PERP | 64065 | 1 | 1 | 18 | |
| TCGA | PERP | 64065 | 11 | 17 | 68 |
Total number of gains: 50; Total number of losses: 50; Total Number of normals: 388.
Somatic mutations of PERP:
Generating mutation plots.
Highly correlated genes for PERP:
Showing top 20/767 corelated genes with mean PCC>0.5.
| Gene1 | Gene2 | Mean PCC | Num. Datasets | Num. PCC<0 | Num. PCC>0.5 |
|---|---|---|---|---|---|
| PERP | DSP | 0.772785 | 12 | 0 | 12 |
| PERP | KRT6C | 0.770313 | 4 | 0 | 4 |
| PERP | DSG3 | 0.757732 | 12 | 0 | 12 |
| PERP | MAL2 | 0.736788 | 7 | 0 | 6 |
| PERP | SLC38A9 | 0.731438 | 3 | 0 | 3 |
| PERP | DSC2 | 0.724663 | 12 | 0 | 11 |
| PERP | ESRP1 | 0.706226 | 10 | 0 | 9 |
| PERP | C6orf132 | 0.702491 | 6 | 0 | 5 |
| PERP | GPR45 | 0.697302 | 4 | 0 | 4 |
| PERP | GRHL3 | 0.697147 | 7 | 0 | 7 |
| PERP | SLC4A9 | 0.692842 | 3 | 0 | 3 |
| PERP | MREG | 0.690712 | 9 | 0 | 9 |
| PERP | CDS1 | 0.689084 | 11 | 0 | 10 |
| PERP | ZNF750 | 0.685065 | 12 | 0 | 10 |
| PERP | S100A11 | 0.683449 | 9 | 0 | 7 |
| PERP | CDH1 | 0.681745 | 11 | 0 | 11 |
| PERP | SERPINB5 | 0.681421 | 11 | 0 | 10 |
| PERP | IL1RN | 0.681398 | 9 | 0 | 7 |
| PERP | SPRR2A | 0.681189 | 4 | 0 | 3 |
| PERP | TMPRSS11E | 0.68073 | 9 | 0 | 8 |
For details and further investigation, click here