| Full name: plexin A3 | Alias Symbol: SEX|XAP-6|6.3|Plxn3 | ||
| Type: protein-coding gene | Cytoband: Xq28 | ||
| Entrez ID: 55558 | HGNC ID: HGNC:9101 | Ensembl Gene: ENSG00000130827 | OMIM ID: 300022 |
PLXNA3 involved pathways:
| KEGG pathway | Description | View |
|---|---|---|
| hsa04360 | Axon guidance |
Expression of PLXNA3:
| Dataset | Gene | EntrezID | Probe | Log2FC | Adj.pValue | Expression |
|---|---|---|---|---|---|---|
| GSE17351 | PLXNA3 | 55558 | 203623_at | 0.4243 | 0.4702 | |
| GSE20347 | PLXNA3 | 55558 | 203623_at | 0.4779 | 0.0000 | |
| GSE23400 | PLXNA3 | 55558 | 203623_at | 0.3889 | 0.0000 | |
| GSE26886 | PLXNA3 | 55558 | 203623_at | 0.1366 | 0.4291 | |
| GSE29001 | PLXNA3 | 55558 | 203623_at | 0.1685 | 0.4882 | |
| GSE38129 | PLXNA3 | 55558 | 203623_at | 0.4343 | 0.0002 | |
| GSE45670 | PLXNA3 | 55558 | 203623_at | 0.5548 | 0.0079 | |
| GSE53622 | PLXNA3 | 55558 | 34159 | 0.7552 | 0.0000 | |
| GSE53624 | PLXNA3 | 55558 | 34159 | 0.1850 | 0.0036 | |
| GSE63941 | PLXNA3 | 55558 | 203623_at | -0.9547 | 0.0122 | |
| GSE77861 | PLXNA3 | 55558 | 203623_at | -0.0243 | 0.8771 | |
| GSE97050 | PLXNA3 | 55558 | A_33_P3413048 | 0.1084 | 0.7290 | |
| SRP007169 | PLXNA3 | 55558 | RNAseq | 1.3039 | 0.0019 | |
| SRP008496 | PLXNA3 | 55558 | RNAseq | 1.9033 | 0.0000 | |
| SRP064894 | PLXNA3 | 55558 | RNAseq | 1.1089 | 0.0000 | |
| SRP133303 | PLXNA3 | 55558 | RNAseq | 0.9945 | 0.0001 | |
| SRP159526 | PLXNA3 | 55558 | RNAseq | 0.5912 | 0.0375 | |
| SRP193095 | PLXNA3 | 55558 | RNAseq | 0.6476 | 0.0000 | |
| SRP219564 | PLXNA3 | 55558 | RNAseq | 0.1746 | 0.6836 | |
| TCGA | PLXNA3 | 55558 | RNAseq | 0.1499 | 0.0292 |
Upregulated datasets: 3; Downregulated datasets: 0.
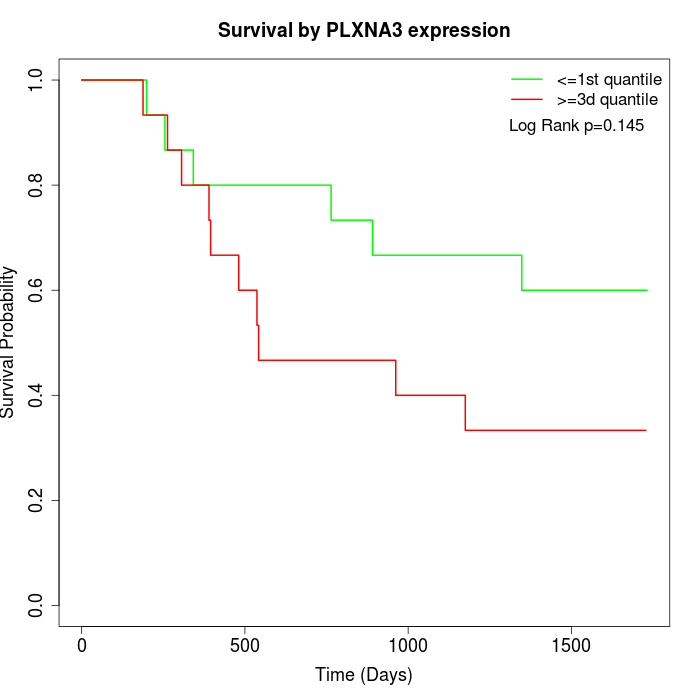
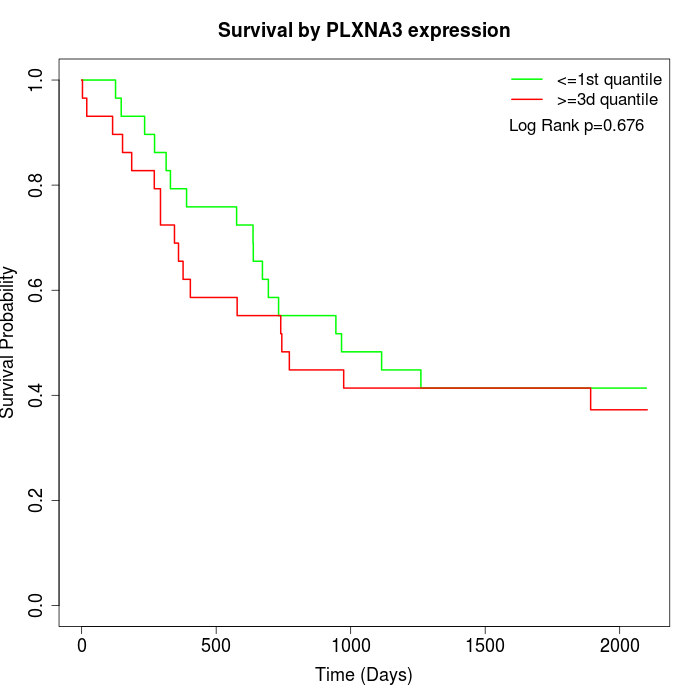
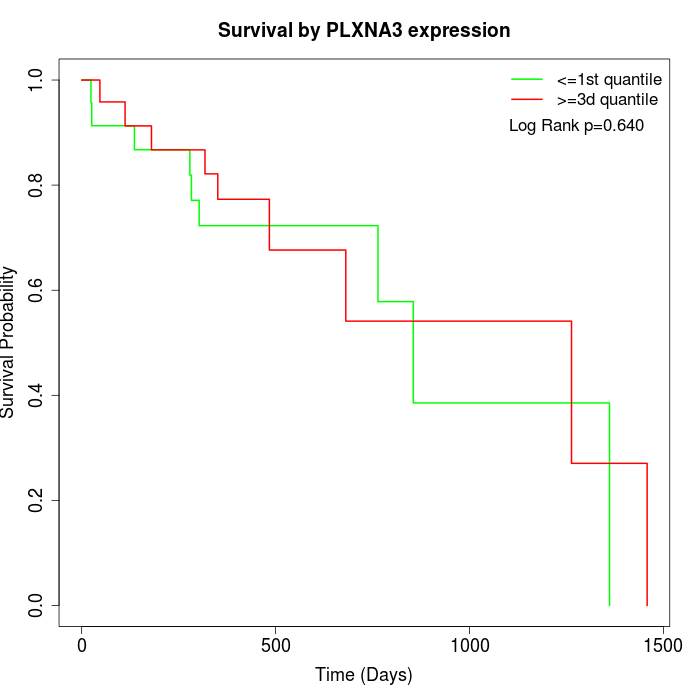
Survival by PLXNA3 expression:
Note: Click image to view full size file.
Copy number change of PLXNA3:
No record found for this gene.
Somatic mutations of PLXNA3:
Generating mutation plots.
Highly correlated genes for PLXNA3:
Showing top 20/171 corelated genes with mean PCC>0.5.
| Gene1 | Gene2 | Mean PCC | Num. Datasets | Num. PCC<0 | Num. PCC>0.5 |
|---|---|---|---|---|---|
| PLXNA3 | HTRA4 | 0.730851 | 3 | 0 | 3 |
| PLXNA3 | FBLN7 | 0.694301 | 4 | 0 | 3 |
| PLXNA3 | KLHDC4 | 0.694187 | 4 | 0 | 3 |
| PLXNA3 | ANKRD33B | 0.68671 | 3 | 0 | 3 |
| PLXNA3 | CLSTN3 | 0.67342 | 4 | 0 | 4 |
| PLXNA3 | EGR4 | 0.670282 | 4 | 0 | 4 |
| PLXNA3 | PARP10 | 0.669661 | 4 | 0 | 4 |
| PLXNA3 | ANXA10 | 0.665141 | 3 | 0 | 3 |
| PLXNA3 | MAP4K1 | 0.660995 | 3 | 0 | 3 |
| PLXNA3 | PGP | 0.654474 | 4 | 0 | 3 |
| PLXNA3 | SCN9A | 0.651787 | 4 | 0 | 3 |
| PLXNA3 | DOHH | 0.647882 | 4 | 0 | 3 |
| PLXNA3 | PSMB11 | 0.645981 | 3 | 0 | 3 |
| PLXNA3 | ZNF576 | 0.644284 | 3 | 0 | 3 |
| PLXNA3 | ARVCF | 0.640977 | 4 | 0 | 3 |
| PLXNA3 | PLEKHG4B | 0.639507 | 3 | 0 | 3 |
| PLXNA3 | YEATS2 | 0.63125 | 7 | 0 | 7 |
| PLXNA3 | CD70 | 0.631071 | 4 | 0 | 4 |
| PLXNA3 | SHROOM1 | 0.629983 | 4 | 0 | 3 |
| PLXNA3 | INF2 | 0.629005 | 4 | 0 | 3 |
For details and further investigation, click here