| Full name: coiled-coil alpha-helical rod protein 1 | Alias Symbol: HCR | ||
| Type: protein-coding gene | Cytoband: 6p21.33 | ||
| Entrez ID: 54535 | HGNC ID: HGNC:13930 | Ensembl Gene: ENSG00000204536 | OMIM ID: 605310 |
Expression of CCHCR1:
| Dataset | Gene | EntrezID | Probe | Log2FC | Adj.pValue | Expression |
|---|---|---|---|---|---|---|
| GSE17351 | CCHCR1 | 54535 | 209698_at | 0.4743 | 0.1533 | |
| GSE20347 | CCHCR1 | 54535 | 209698_at | 0.2789 | 0.0085 | |
| GSE23400 | CCHCR1 | 54535 | 209698_at | 0.1370 | 0.0020 | |
| GSE26886 | CCHCR1 | 54535 | 37424_at | -0.0505 | 0.7677 | |
| GSE29001 | CCHCR1 | 54535 | 209698_at | 0.3568 | 0.0208 | |
| GSE38129 | CCHCR1 | 54535 | 209698_at | 0.2847 | 0.0006 | |
| GSE45670 | CCHCR1 | 54535 | 209698_at | 0.4587 | 0.0006 | |
| GSE53622 | CCHCR1 | 54535 | 13250 | 0.0608 | 0.3387 | |
| GSE53624 | CCHCR1 | 54535 | 13250 | 0.2542 | 0.0000 | |
| GSE63941 | CCHCR1 | 54535 | 209698_at | 0.8025 | 0.0146 | |
| GSE77861 | CCHCR1 | 54535 | 209698_at | 0.0486 | 0.7393 | |
| GSE97050 | CCHCR1 | 54535 | A_23_P145330 | 0.1767 | 0.4988 | |
| SRP007169 | CCHCR1 | 54535 | RNAseq | 0.4575 | 0.3931 | |
| SRP008496 | CCHCR1 | 54535 | RNAseq | 0.2852 | 0.4607 | |
| SRP064894 | CCHCR1 | 54535 | RNAseq | 1.1791 | 0.0000 | |
| SRP133303 | CCHCR1 | 54535 | RNAseq | 0.2236 | 0.2597 | |
| SRP159526 | CCHCR1 | 54535 | RNAseq | 0.9892 | 0.0745 | |
| SRP193095 | CCHCR1 | 54535 | RNAseq | 0.3051 | 0.0101 | |
| SRP219564 | CCHCR1 | 54535 | RNAseq | 0.3986 | 0.3112 | |
| TCGA | CCHCR1 | 54535 | RNAseq | 0.0797 | 0.2177 |
Upregulated datasets: 1; Downregulated datasets: 0.
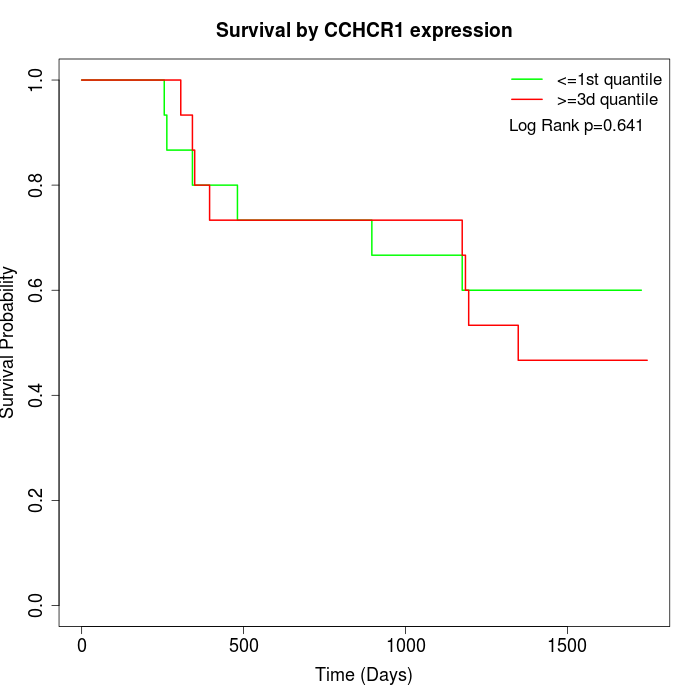
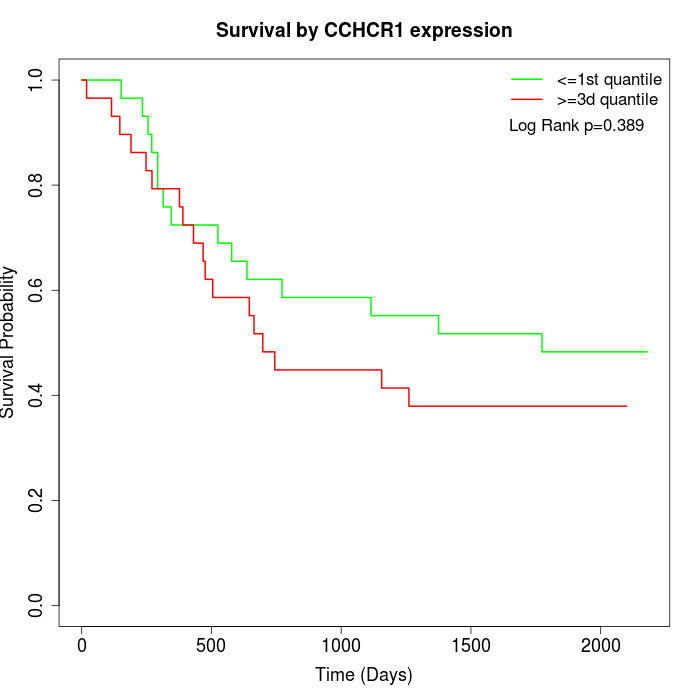
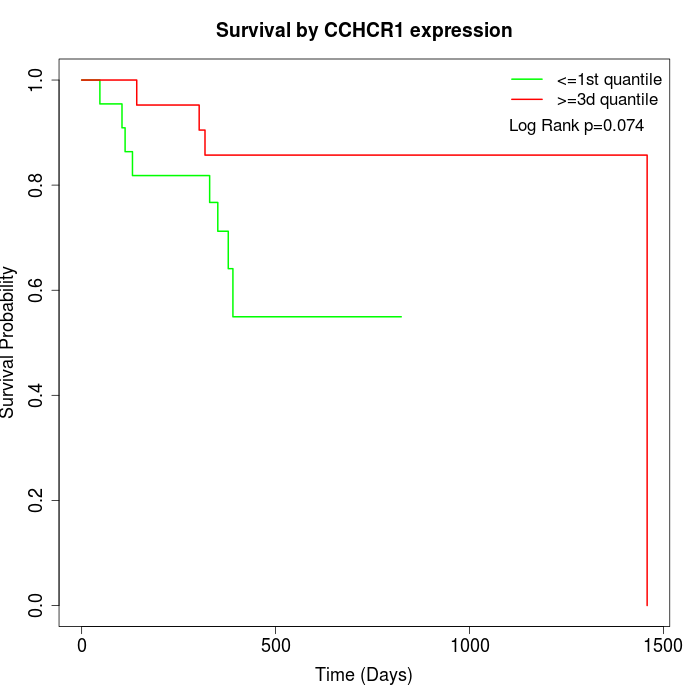
Survival by CCHCR1 expression:
Note: Click image to view full size file.
Copy number change of CCHCR1:
| Dataset | Gene | EntrezID | Gain | Loss | Normal | Detail |
|---|---|---|---|---|---|---|
| GSE15526 | CCHCR1 | 54535 | 5 | 1 | 24 | |
| GSE20123 | CCHCR1 | 54535 | 5 | 1 | 24 | |
| GSE43470 | CCHCR1 | 54535 | 5 | 1 | 37 | |
| GSE46452 | CCHCR1 | 54535 | 1 | 10 | 48 | |
| GSE47630 | CCHCR1 | 54535 | 7 | 3 | 30 | |
| GSE54993 | CCHCR1 | 54535 | 2 | 1 | 67 | |
| GSE54994 | CCHCR1 | 54535 | 11 | 4 | 38 | |
| GSE60625 | CCHCR1 | 54535 | 0 | 1 | 10 | |
| GSE74703 | CCHCR1 | 54535 | 5 | 0 | 31 | |
| GSE74704 | CCHCR1 | 54535 | 2 | 0 | 18 | |
| TCGA | CCHCR1 | 54535 | 17 | 16 | 63 |
Total number of gains: 60; Total number of losses: 38; Total Number of normals: 390.
Somatic mutations of CCHCR1:
Generating mutation plots.
Highly correlated genes for CCHCR1:
Showing top 20/247 corelated genes with mean PCC>0.5.
| Gene1 | Gene2 | Mean PCC | Num. Datasets | Num. PCC<0 | Num. PCC>0.5 |
|---|---|---|---|---|---|
| CCHCR1 | ISG15 | 0.692147 | 3 | 0 | 3 |
| CCHCR1 | LDB1 | 0.681608 | 4 | 0 | 4 |
| CCHCR1 | BCL9 | 0.653242 | 3 | 0 | 3 |
| CCHCR1 | TRIM9 | 0.648111 | 4 | 0 | 3 |
| CCHCR1 | USP18 | 0.638903 | 4 | 0 | 3 |
| CCHCR1 | PADI2 | 0.634811 | 5 | 0 | 4 |
| CCHCR1 | NADK | 0.634027 | 5 | 0 | 5 |
| CCHCR1 | COMMD7 | 0.633982 | 3 | 0 | 3 |
| CCHCR1 | PFN1 | 0.633523 | 4 | 0 | 3 |
| CCHCR1 | IFI6 | 0.626283 | 4 | 0 | 3 |
| CCHCR1 | ZNF496 | 0.6261 | 3 | 0 | 3 |
| CCHCR1 | HLA-F | 0.620825 | 4 | 0 | 3 |
| CCHCR1 | UBE2T | 0.620464 | 3 | 0 | 3 |
| CCHCR1 | APOL1 | 0.620209 | 3 | 0 | 3 |
| CCHCR1 | CXCL11 | 0.619553 | 3 | 0 | 3 |
| CCHCR1 | ELK4 | 0.613698 | 4 | 0 | 3 |
| CCHCR1 | ASB2 | 0.613439 | 3 | 0 | 3 |
| CCHCR1 | IRF9 | 0.613417 | 5 | 0 | 4 |
| CCHCR1 | PPP1R18 | 0.611068 | 3 | 0 | 3 |
| CCHCR1 | IFI35 | 0.608989 | 4 | 0 | 3 |
For details and further investigation, click here