| Full name: CKLF like MARVEL transmembrane domain containing 2 | Alias Symbol: MGC39436|FLJ25732 | ||
| Type: protein-coding gene | Cytoband: 16q21 | ||
| Entrez ID: 146225 | HGNC ID: HGNC:19173 | Ensembl Gene: ENSG00000140932 | OMIM ID: 607885 |
Expression of CMTM2:
| Dataset | Gene | EntrezID | Probe | Log2FC | Adj.pValue | Expression |
|---|---|---|---|---|---|---|
| GSE17351 | CMTM2 | 146225 | 229967_at | 0.6577 | 0.3333 | |
| GSE26886 | CMTM2 | 146225 | 229967_at | 0.1077 | 0.3772 | |
| GSE45670 | CMTM2 | 146225 | 229967_at | 0.2828 | 0.1200 | |
| GSE53622 | CMTM2 | 146225 | 10259 | 0.3664 | 0.0719 | |
| GSE53624 | CMTM2 | 146225 | 10259 | 0.4583 | 0.0414 | |
| GSE63941 | CMTM2 | 146225 | 229967_at | 0.1133 | 0.5358 | |
| GSE77861 | CMTM2 | 146225 | 229967_at | -0.0666 | 0.3866 | |
| GSE97050 | CMTM2 | 146225 | A_23_P152234 | -0.1517 | 0.6629 | |
| SRP159526 | CMTM2 | 146225 | RNAseq | 3.8427 | 0.0001 | |
| TCGA | CMTM2 | 146225 | RNAseq | -1.1192 | 0.0022 |
Upregulated datasets: 1; Downregulated datasets: 1.
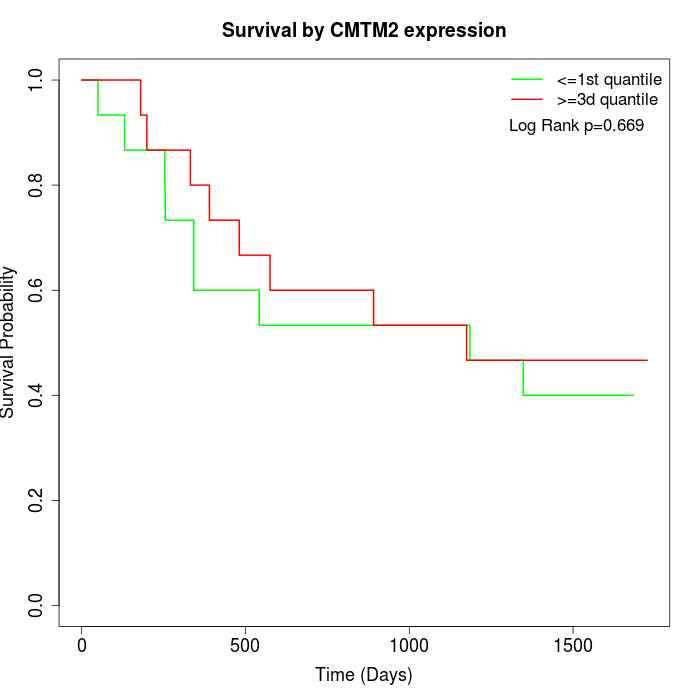
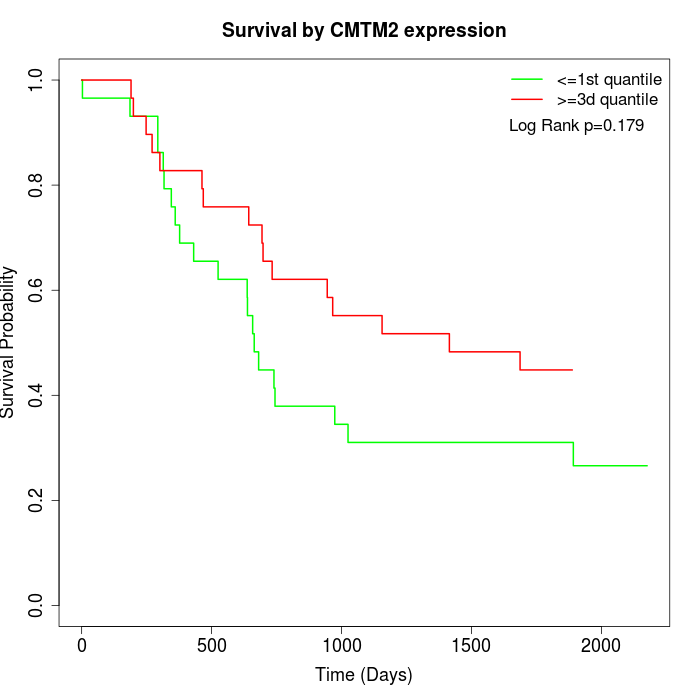
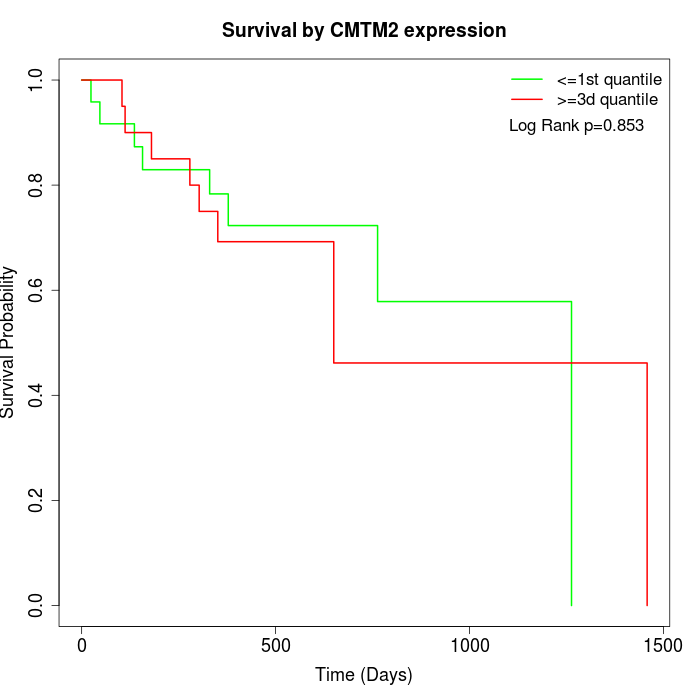
Survival by CMTM2 expression:
Note: Click image to view full size file.
Copy number change of CMTM2:
| Dataset | Gene | EntrezID | Gain | Loss | Normal | Detail |
|---|---|---|---|---|---|---|
| GSE15526 | CMTM2 | 146225 | 8 | 2 | 20 | |
| GSE20123 | CMTM2 | 146225 | 8 | 2 | 20 | |
| GSE43470 | CMTM2 | 146225 | 1 | 8 | 34 | |
| GSE46452 | CMTM2 | 146225 | 38 | 1 | 20 | |
| GSE47630 | CMTM2 | 146225 | 10 | 8 | 22 | |
| GSE54993 | CMTM2 | 146225 | 2 | 4 | 64 | |
| GSE54994 | CMTM2 | 146225 | 8 | 10 | 35 | |
| GSE60625 | CMTM2 | 146225 | 4 | 0 | 7 | |
| GSE74703 | CMTM2 | 146225 | 1 | 5 | 30 | |
| GSE74704 | CMTM2 | 146225 | 5 | 1 | 14 | |
| TCGA | CMTM2 | 146225 | 30 | 11 | 55 |
Total number of gains: 115; Total number of losses: 52; Total Number of normals: 321.
Somatic mutations of CMTM2:
Generating mutation plots.
Highly correlated genes for CMTM2:
Showing top 20/54 corelated genes with mean PCC>0.5.
| Gene1 | Gene2 | Mean PCC | Num. Datasets | Num. PCC<0 | Num. PCC>0.5 |
|---|---|---|---|---|---|
| CMTM2 | PROK2 | 0.825001 | 5 | 0 | 5 |
| CMTM2 | AQP9 | 0.788005 | 5 | 0 | 5 |
| CMTM2 | FPR1 | 0.75126 | 4 | 0 | 4 |
| CMTM2 | CXCR1 | 0.745265 | 5 | 0 | 4 |
| CMTM2 | PF4V1 | 0.708505 | 3 | 0 | 3 |
| CMTM2 | C20orf202 | 0.683014 | 3 | 0 | 3 |
| CMTM2 | PADI4 | 0.668973 | 4 | 0 | 3 |
| CMTM2 | LILRA6 | 0.65858 | 4 | 0 | 4 |
| CMTM2 | CA3 | 0.656603 | 3 | 0 | 3 |
| CMTM2 | MGAM | 0.653957 | 6 | 0 | 5 |
| CMTM2 | CSF3R | 0.651878 | 4 | 0 | 3 |
| CMTM2 | NCAM1 | 0.645452 | 3 | 0 | 3 |
| CMTM2 | GLT1D1 | 0.641542 | 4 | 0 | 3 |
| CMTM2 | LILRA3 | 0.639027 | 3 | 0 | 3 |
| CMTM2 | DACT2 | 0.635389 | 3 | 0 | 3 |
| CMTM2 | MNDA | 0.631264 | 6 | 0 | 5 |
| CMTM2 | IL10 | 0.631108 | 4 | 0 | 3 |
| CMTM2 | CLEC4A | 0.626983 | 4 | 0 | 4 |
| CMTM2 | G0S2 | 0.61852 | 5 | 0 | 4 |
| CMTM2 | FCAR | 0.61625 | 4 | 0 | 4 |
For details and further investigation, click here