| Full name: LDL receptor related protein 1 | Alias Symbol: LRP|CD91|LRP1A|APOER | ||
| Type: protein-coding gene | Cytoband: 12q13.3 | ||
| Entrez ID: 4035 | HGNC ID: HGNC:6692 | Ensembl Gene: ENSG00000123384 | OMIM ID: 107770 |
LRP1 involved pathways:
| KEGG pathway | Description | View |
|---|---|---|
| hsa05010 | Alzheimer's disease |
Expression of LRP1:
| Dataset | Gene | EntrezID | Probe | Log2FC | Adj.pValue | Expression |
|---|---|---|---|---|---|---|
| GSE17351 | LRP1 | 4035 | 200785_s_at | 0.1225 | 0.6992 | |
| GSE20347 | LRP1 | 4035 | 200784_s_at | 0.0211 | 0.8111 | |
| GSE23400 | LRP1 | 4035 | 200785_s_at | 0.1279 | 0.0855 | |
| GSE26886 | LRP1 | 4035 | 200784_s_at | 0.4870 | 0.0157 | |
| GSE29001 | LRP1 | 4035 | 200785_s_at | 0.0925 | 0.7332 | |
| GSE38129 | LRP1 | 4035 | 200784_s_at | -0.0650 | 0.4902 | |
| GSE45670 | LRP1 | 4035 | 200784_s_at | -0.0133 | 0.9334 | |
| GSE53622 | LRP1 | 4035 | 133921 | 0.2281 | 0.0208 | |
| GSE53624 | LRP1 | 4035 | 64670 | -0.2008 | 0.0075 | |
| GSE63941 | LRP1 | 4035 | 200784_s_at | -0.2761 | 0.2244 | |
| GSE77861 | LRP1 | 4035 | 200785_s_at | 0.1399 | 0.5539 | |
| GSE97050 | LRP1 | 4035 | A_23_P124837 | 0.4783 | 0.2411 | |
| SRP007169 | LRP1 | 4035 | RNAseq | 1.9553 | 0.0000 | |
| SRP008496 | LRP1 | 4035 | RNAseq | 2.1454 | 0.0000 | |
| SRP064894 | LRP1 | 4035 | RNAseq | 0.5397 | 0.0287 | |
| SRP133303 | LRP1 | 4035 | RNAseq | 0.1163 | 0.6744 | |
| SRP159526 | LRP1 | 4035 | RNAseq | 0.1065 | 0.7142 | |
| SRP193095 | LRP1 | 4035 | RNAseq | 0.6727 | 0.0000 | |
| SRP219564 | LRP1 | 4035 | RNAseq | 0.1189 | 0.7960 | |
| TCGA | LRP1 | 4035 | RNAseq | -0.0736 | 0.2577 |
Upregulated datasets: 2; Downregulated datasets: 0.
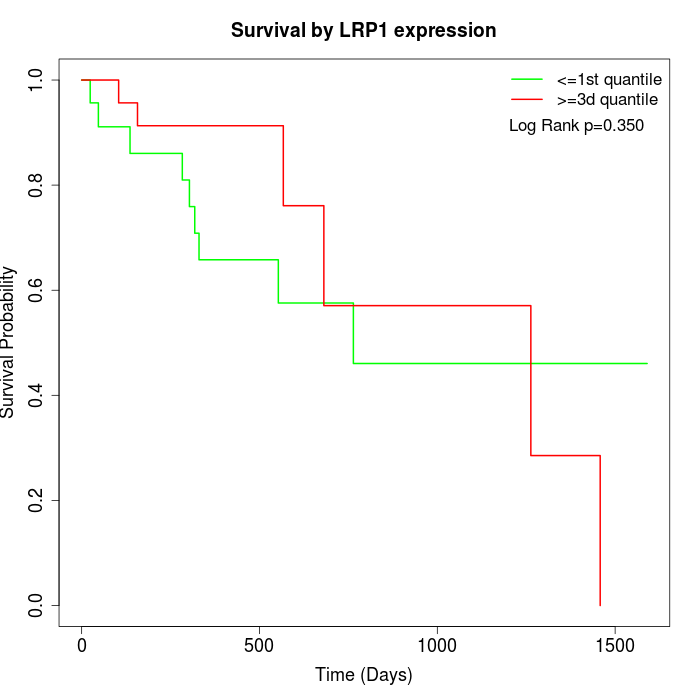
Survival by LRP1 expression:
Note: Click image to view full size file.
Copy number change of LRP1:
| Dataset | Gene | EntrezID | Gain | Loss | Normal | Detail |
|---|---|---|---|---|---|---|
| GSE15526 | LRP1 | 4035 | 7 | 1 | 22 | |
| GSE20123 | LRP1 | 4035 | 7 | 0 | 23 | |
| GSE43470 | LRP1 | 4035 | 3 | 0 | 40 | |
| GSE46452 | LRP1 | 4035 | 8 | 1 | 50 | |
| GSE47630 | LRP1 | 4035 | 10 | 2 | 28 | |
| GSE54993 | LRP1 | 4035 | 0 | 5 | 65 | |
| GSE54994 | LRP1 | 4035 | 4 | 1 | 48 | |
| GSE60625 | LRP1 | 4035 | 0 | 0 | 11 | |
| GSE74703 | LRP1 | 4035 | 3 | 0 | 33 | |
| GSE74704 | LRP1 | 4035 | 4 | 0 | 16 | |
| TCGA | LRP1 | 4035 | 14 | 10 | 72 |
Total number of gains: 60; Total number of losses: 20; Total Number of normals: 408.
Somatic mutations of LRP1:
Generating mutation plots.
Highly correlated genes for LRP1:
Showing top 20/515 corelated genes with mean PCC>0.5.
| Gene1 | Gene2 | Mean PCC | Num. Datasets | Num. PCC<0 | Num. PCC>0.5 |
|---|---|---|---|---|---|
| LRP1 | SIX5 | 0.766639 | 3 | 0 | 3 |
| LRP1 | GMPPA | 0.756151 | 3 | 0 | 3 |
| LRP1 | ACIN1 | 0.747823 | 3 | 0 | 3 |
| LRP1 | INTS1 | 0.718306 | 3 | 0 | 3 |
| LRP1 | LNP1 | 0.71322 | 3 | 0 | 3 |
| LRP1 | SOX17 | 0.707202 | 3 | 0 | 3 |
| LRP1 | MYPOP | 0.689883 | 4 | 0 | 3 |
| LRP1 | TNFRSF4 | 0.68665 | 3 | 0 | 3 |
| LRP1 | ZC3H3 | 0.684153 | 3 | 0 | 3 |
| LRP1 | BCAS4 | 0.683506 | 3 | 0 | 3 |
| LRP1 | ADRM1 | 0.68016 | 3 | 0 | 3 |
| LRP1 | MKNK1 | 0.676143 | 3 | 0 | 3 |
| LRP1 | ARF3 | 0.674826 | 3 | 0 | 3 |
| LRP1 | PACS1 | 0.672671 | 4 | 0 | 3 |
| LRP1 | SYNGR2 | 0.67064 | 3 | 0 | 3 |
| LRP1 | PRR12 | 0.670114 | 3 | 0 | 3 |
| LRP1 | EEFSEC | 0.66721 | 4 | 0 | 4 |
| LRP1 | IMPDH1 | 0.665847 | 4 | 0 | 4 |
| LRP1 | RBM14 | 0.665239 | 4 | 0 | 4 |
| LRP1 | SIK1 | 0.662958 | 3 | 0 | 3 |
For details and further investigation, click here