| Full name: lumican | Alias Symbol: SLRR2D | ||
| Type: protein-coding gene | Cytoband: 12q21.33 | ||
| Entrez ID: 4060 | HGNC ID: HGNC:6724 | Ensembl Gene: ENSG00000139329 | OMIM ID: 600616 |
| Related drugs: RIZATRIPTAN... [more] | |||
LUM involved pathways:
| KEGG pathway | Description | View |
|---|---|---|
| hsa05205 | Proteoglycans in cancer |
Expression of LUM:
| Dataset | Gene | EntrezID | Probe | Log2FC | Adj.pValue | Expression |
|---|---|---|---|---|---|---|
| GSE17351 | LUM | 4060 | 201744_s_at | 0.3693 | 0.6462 | |
| GSE20347 | LUM | 4060 | 201744_s_at | 2.4229 | 0.0000 | |
| GSE23400 | LUM | 4060 | 201744_s_at | 1.4557 | 0.0000 | |
| GSE26886 | LUM | 4060 | 201744_s_at | 2.8540 | 0.0001 | |
| GSE29001 | LUM | 4060 | 201744_s_at | 1.6643 | 0.0164 | |
| GSE38129 | LUM | 4060 | 201744_s_at | 1.6618 | 0.0000 | |
| GSE45670 | LUM | 4060 | 201744_s_at | 0.0079 | 0.9903 | |
| GSE53622 | LUM | 4060 | 81306 | 1.8627 | 0.0000 | |
| GSE53624 | LUM | 4060 | 81306 | 2.1594 | 0.0000 | |
| GSE63941 | LUM | 4060 | 201744_s_at | -8.7738 | 0.0001 | |
| GSE77861 | LUM | 4060 | 201744_s_at | 3.6108 | 0.0020 | |
| GSE97050 | LUM | 4060 | A_23_P99063 | 1.7075 | 0.1296 | |
| SRP007169 | LUM | 4060 | RNAseq | 4.5536 | 0.0000 | |
| SRP008496 | LUM | 4060 | RNAseq | 4.9412 | 0.0000 | |
| SRP064894 | LUM | 4060 | RNAseq | 1.6462 | 0.0000 | |
| SRP133303 | LUM | 4060 | RNAseq | 2.1813 | 0.0000 | |
| SRP159526 | LUM | 4060 | RNAseq | 1.9021 | 0.0003 | |
| SRP193095 | LUM | 4060 | RNAseq | 3.2397 | 0.0000 | |
| SRP219564 | LUM | 4060 | RNAseq | 1.7064 | 0.0231 | |
| TCGA | LUM | 4060 | RNAseq | 0.2710 | 0.0062 |
Upregulated datasets: 15; Downregulated datasets: 1.
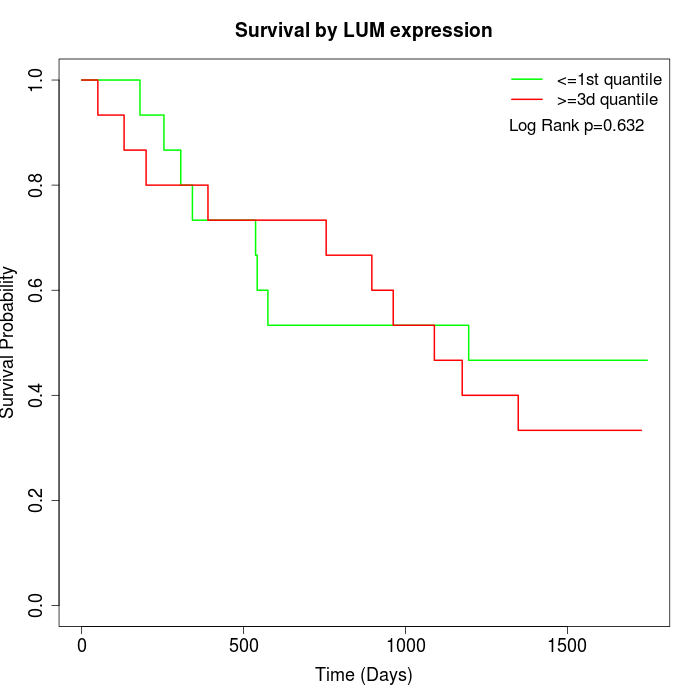
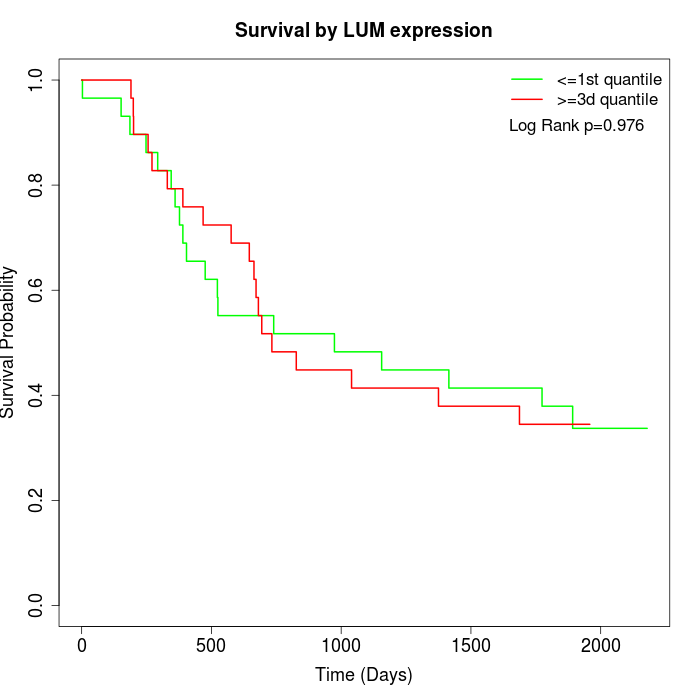
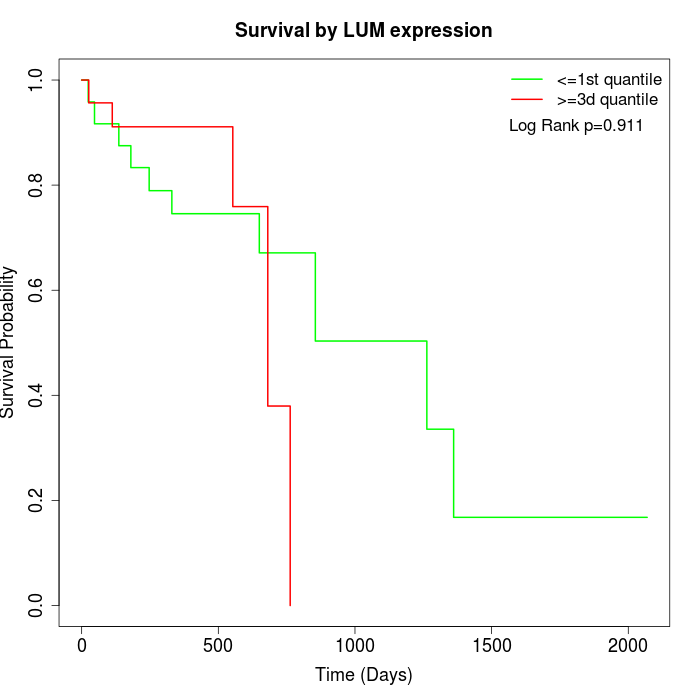
Survival by LUM expression:
Note: Click image to view full size file.
Copy number change of LUM:
| Dataset | Gene | EntrezID | Gain | Loss | Normal | Detail |
|---|---|---|---|---|---|---|
| GSE15526 | LUM | 4060 | 3 | 2 | 25 | |
| GSE20123 | LUM | 4060 | 3 | 2 | 25 | |
| GSE43470 | LUM | 4060 | 3 | 0 | 40 | |
| GSE46452 | LUM | 4060 | 9 | 1 | 49 | |
| GSE47630 | LUM | 4060 | 10 | 1 | 29 | |
| GSE54993 | LUM | 4060 | 0 | 6 | 64 | |
| GSE54994 | LUM | 4060 | 6 | 1 | 46 | |
| GSE60625 | LUM | 4060 | 0 | 0 | 11 | |
| GSE74703 | LUM | 4060 | 3 | 0 | 33 | |
| GSE74704 | LUM | 4060 | 1 | 2 | 17 | |
| TCGA | LUM | 4060 | 17 | 12 | 67 |
Total number of gains: 55; Total number of losses: 27; Total Number of normals: 406.
Somatic mutations of LUM:
Generating mutation plots.
Highly correlated genes for LUM:
Showing top 20/899 corelated genes with mean PCC>0.5.
| Gene1 | Gene2 | Mean PCC | Num. Datasets | Num. PCC<0 | Num. PCC>0.5 |
|---|---|---|---|---|---|
| LUM | COL6A3 | 0.850557 | 13 | 0 | 13 |
| LUM | COL3A1 | 0.837484 | 13 | 0 | 13 |
| LUM | CTSK | 0.830216 | 13 | 0 | 13 |
| LUM | VCAN | 0.828979 | 13 | 0 | 13 |
| LUM | SPARC | 0.816119 | 13 | 0 | 13 |
| LUM | COL12A1 | 0.811025 | 9 | 0 | 9 |
| LUM | COL1A2 | 0.80994 | 13 | 0 | 12 |
| LUM | COL5A2 | 0.794408 | 13 | 0 | 13 |
| LUM | FBN1 | 0.789089 | 10 | 0 | 10 |
| LUM | FN1 | 0.784572 | 12 | 0 | 12 |
| LUM | SULF1 | 0.780746 | 13 | 0 | 13 |
| LUM | FNDC1 | 0.773489 | 8 | 0 | 8 |
| LUM | POSTN | 0.772142 | 13 | 0 | 13 |
| LUM | HTRA1 | 0.771013 | 13 | 0 | 13 |
| LUM | CDH11 | 0.76018 | 13 | 0 | 13 |
| LUM | COL1A1 | 0.753344 | 12 | 0 | 12 |
| LUM | MMP2 | 0.750876 | 12 | 0 | 11 |
| LUM | LGALS1 | 0.737922 | 12 | 0 | 11 |
| LUM | FSTL1 | 0.735333 | 13 | 0 | 13 |
| LUM | COL5A1 | 0.734245 | 13 | 0 | 13 |
For details and further investigation, click here