| Full name: microtubule affinity regulating kinase 2 | Alias Symbol: PAR-1|Par1b|PAR-1B | ||
| Type: protein-coding gene | Cytoband: 11q13.1 | ||
| Entrez ID: 2011 | HGNC ID: HGNC:3332 | Ensembl Gene: ENSG00000072518 | OMIM ID: 600526 |
| Related drugs: ACETAMINOPHEN... [more] | |||
Screen Evidence:
| |||
Expression of MARK2:
| Dataset | Gene | EntrezID | Probe | Log2FC | Adj.pValue | Expression |
|---|---|---|---|---|---|---|
| GSE17351 | MARK2 | 2011 | 203942_s_at | 0.1126 | 0.8641 | |
| GSE20347 | MARK2 | 2011 | 203942_s_at | -0.5870 | 0.0004 | |
| GSE23400 | MARK2 | 2011 | 203942_s_at | -0.3926 | 0.0007 | |
| GSE26886 | MARK2 | 2011 | 203942_s_at | -0.3450 | 0.1231 | |
| GSE29001 | MARK2 | 2011 | 203942_s_at | -0.8590 | 0.0070 | |
| GSE38129 | MARK2 | 2011 | 203942_s_at | -0.3167 | 0.0380 | |
| GSE45670 | MARK2 | 2011 | 203942_s_at | 0.0111 | 0.9566 | |
| GSE53622 | MARK2 | 2011 | 80466 | -0.5412 | 0.0000 | |
| GSE53624 | MARK2 | 2011 | 23156 | -0.4137 | 0.0000 | |
| GSE63941 | MARK2 | 2011 | 203942_s_at | 1.1764 | 0.0125 | |
| GSE77861 | MARK2 | 2011 | 203942_s_at | 0.0506 | 0.8289 | |
| GSE97050 | MARK2 | 2011 | A_24_P914495 | 0.1384 | 0.8164 | |
| SRP007169 | MARK2 | 2011 | RNAseq | -1.5976 | 0.0000 | |
| SRP008496 | MARK2 | 2011 | RNAseq | -1.6018 | 0.0000 | |
| SRP064894 | MARK2 | 2011 | RNAseq | -0.8853 | 0.0000 | |
| SRP133303 | MARK2 | 2011 | RNAseq | -0.6221 | 0.0015 | |
| SRP159526 | MARK2 | 2011 | RNAseq | -0.5841 | 0.0008 | |
| SRP193095 | MARK2 | 2011 | RNAseq | -0.4040 | 0.0006 | |
| SRP219564 | MARK2 | 2011 | RNAseq | -0.7225 | 0.0810 | |
| TCGA | MARK2 | 2011 | RNAseq | 0.1135 | 0.0232 |
Upregulated datasets: 1; Downregulated datasets: 2.
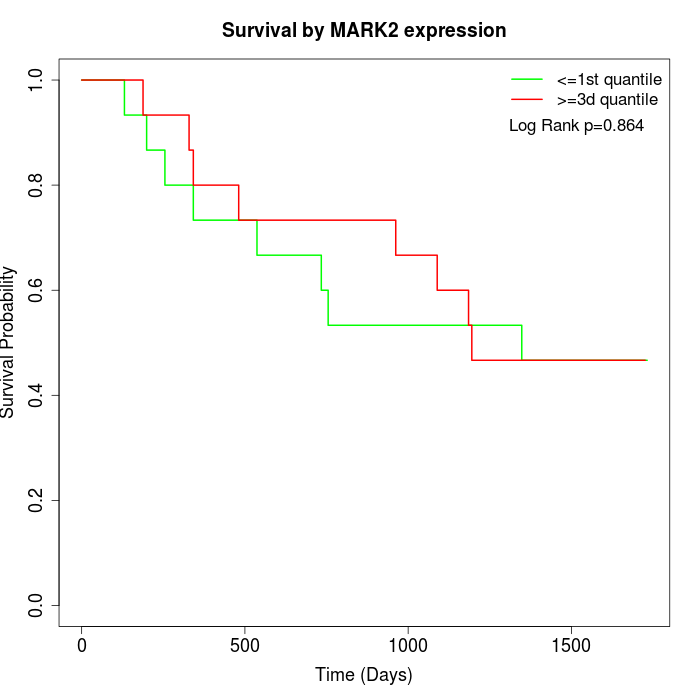
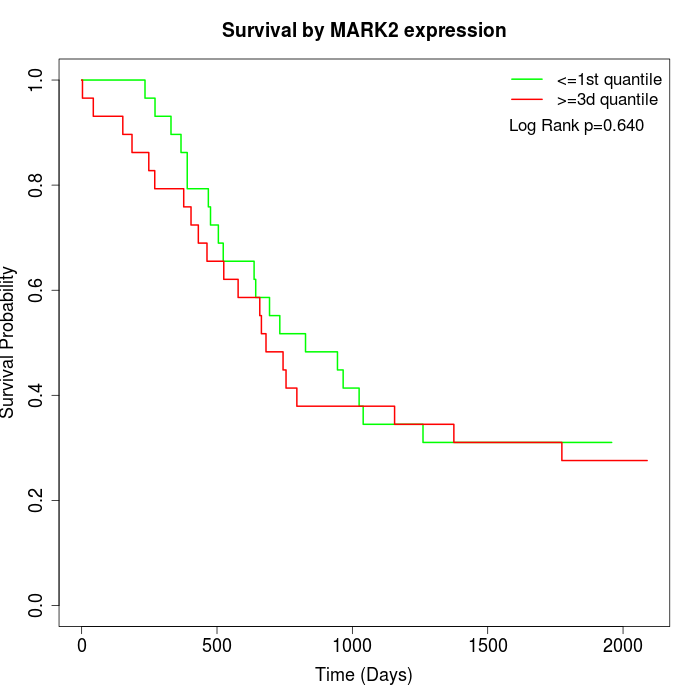
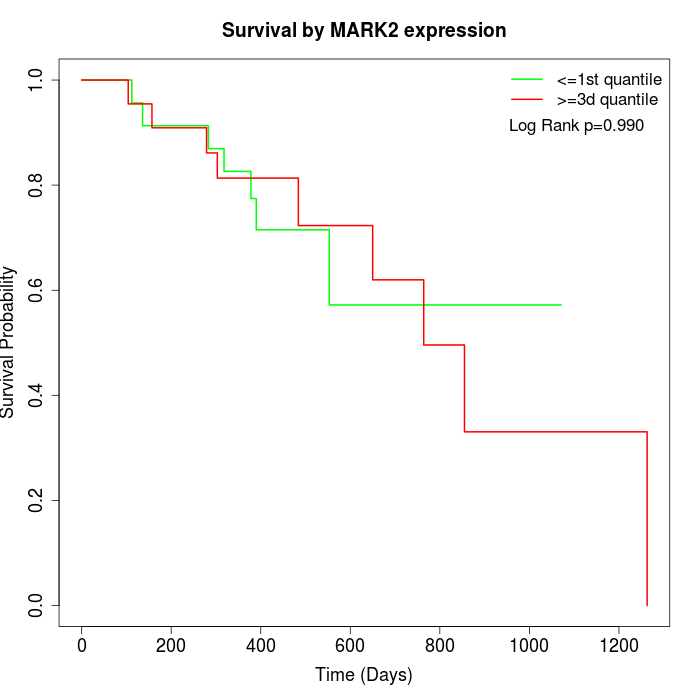
Survival by MARK2 expression:
Note: Click image to view full size file.
Copy number change of MARK2:
| Dataset | Gene | EntrezID | Gain | Loss | Normal | Detail |
|---|---|---|---|---|---|---|
| GSE15526 | MARK2 | 2011 | 6 | 5 | 19 | |
| GSE20123 | MARK2 | 2011 | 6 | 5 | 19 | |
| GSE43470 | MARK2 | 2011 | 1 | 4 | 38 | |
| GSE46452 | MARK2 | 2011 | 8 | 4 | 47 | |
| GSE47630 | MARK2 | 2011 | 3 | 7 | 30 | |
| GSE54993 | MARK2 | 2011 | 3 | 0 | 67 | |
| GSE54994 | MARK2 | 2011 | 5 | 5 | 43 | |
| GSE60625 | MARK2 | 2011 | 0 | 4 | 7 | |
| GSE74703 | MARK2 | 2011 | 1 | 2 | 33 | |
| GSE74704 | MARK2 | 2011 | 4 | 3 | 13 | |
| TCGA | MARK2 | 2011 | 17 | 10 | 69 |
Total number of gains: 54; Total number of losses: 49; Total Number of normals: 385.
Somatic mutations of MARK2:
Generating mutation plots.
Highly correlated genes for MARK2:
Showing top 20/905 corelated genes with mean PCC>0.5.
| Gene1 | Gene2 | Mean PCC | Num. Datasets | Num. PCC<0 | Num. PCC>0.5 |
|---|---|---|---|---|---|
| MARK2 | OTUB1 | 0.827483 | 3 | 0 | 3 |
| MARK2 | DDI2 | 0.778111 | 3 | 0 | 3 |
| MARK2 | EBPL | 0.763697 | 3 | 0 | 3 |
| MARK2 | HEATR5B | 0.749888 | 3 | 0 | 3 |
| MARK2 | RNF149 | 0.734083 | 3 | 0 | 3 |
| MARK2 | PCNT | 0.721395 | 3 | 0 | 3 |
| MARK2 | CBR3-AS1 | 0.702757 | 3 | 0 | 3 |
| MARK2 | CHMP4C | 0.70034 | 5 | 0 | 4 |
| MARK2 | INO80E | 0.698906 | 4 | 0 | 4 |
| MARK2 | RAB3D | 0.698124 | 6 | 0 | 6 |
| MARK2 | TBC1D22B | 0.693773 | 5 | 0 | 4 |
| MARK2 | ADCY7 | 0.693641 | 4 | 0 | 4 |
| MARK2 | FBXL19 | 0.692325 | 4 | 0 | 4 |
| MARK2 | CDS1 | 0.689347 | 8 | 0 | 8 |
| MARK2 | PRKCH | 0.688075 | 9 | 0 | 7 |
| MARK2 | CAPN1 | 0.686198 | 13 | 0 | 10 |
| MARK2 | VPS37B | 0.681781 | 10 | 0 | 10 |
| MARK2 | STIP1 | 0.681395 | 5 | 0 | 4 |
| MARK2 | CERS3 | 0.679928 | 4 | 0 | 4 |
| MARK2 | ARF6 | 0.675241 | 7 | 0 | 6 |
For details and further investigation, click here