| Full name: phosphatidylinositol-5-phosphate 4-kinase type 2 beta | Alias Symbol: PIP5KIIB|PIP5KIIbeta | ||
| Type: protein-coding gene | Cytoband: 17q12 | ||
| Entrez ID: 8396 | HGNC ID: HGNC:8998 | Ensembl Gene: ENSG00000276293 | OMIM ID: 603261 |
PIP4K2B involved pathways:
| KEGG pathway | Description | View |
|---|---|---|
| hsa04810 | Regulation of actin cytoskeleton |
Expression of PIP4K2B:
| Dataset | Gene | EntrezID | Probe | Log2FC | Adj.pValue | Expression |
|---|---|---|---|---|---|---|
| GSE17351 | PIP4K2B | 8396 | 201080_at | 0.2317 | 0.4062 | |
| GSE20347 | PIP4K2B | 8396 | 201080_at | 0.1609 | 0.2648 | |
| GSE23400 | PIP4K2B | 8396 | 201080_at | -0.0431 | 0.4433 | |
| GSE26886 | PIP4K2B | 8396 | 201080_at | 0.4596 | 0.0163 | |
| GSE29001 | PIP4K2B | 8396 | 201080_at | 0.1374 | 0.5010 | |
| GSE38129 | PIP4K2B | 8396 | 201080_at | 0.0147 | 0.9182 | |
| GSE45670 | PIP4K2B | 8396 | 201080_at | -0.0493 | 0.8384 | |
| GSE53622 | PIP4K2B | 8396 | 94067 | 0.0117 | 0.9032 | |
| GSE53624 | PIP4K2B | 8396 | 94067 | 0.1982 | 0.0009 | |
| GSE63941 | PIP4K2B | 8396 | 201080_at | -0.3820 | 0.2737 | |
| GSE77861 | PIP4K2B | 8396 | 201080_at | 0.1681 | 0.5117 | |
| GSE97050 | PIP4K2B | 8396 | A_23_P407115 | 0.0312 | 0.9113 | |
| SRP007169 | PIP4K2B | 8396 | RNAseq | 0.7094 | 0.0541 | |
| SRP008496 | PIP4K2B | 8396 | RNAseq | 0.7807 | 0.0002 | |
| SRP064894 | PIP4K2B | 8396 | RNAseq | 0.4560 | 0.0009 | |
| SRP133303 | PIP4K2B | 8396 | RNAseq | 0.1007 | 0.4677 | |
| SRP159526 | PIP4K2B | 8396 | RNAseq | 0.4837 | 0.1138 | |
| SRP193095 | PIP4K2B | 8396 | RNAseq | 0.3394 | 0.0000 | |
| SRP219564 | PIP4K2B | 8396 | RNAseq | 0.0527 | 0.8739 | |
| TCGA | PIP4K2B | 8396 | RNAseq | -0.0199 | 0.6583 |
Upregulated datasets: 0; Downregulated datasets: 0.
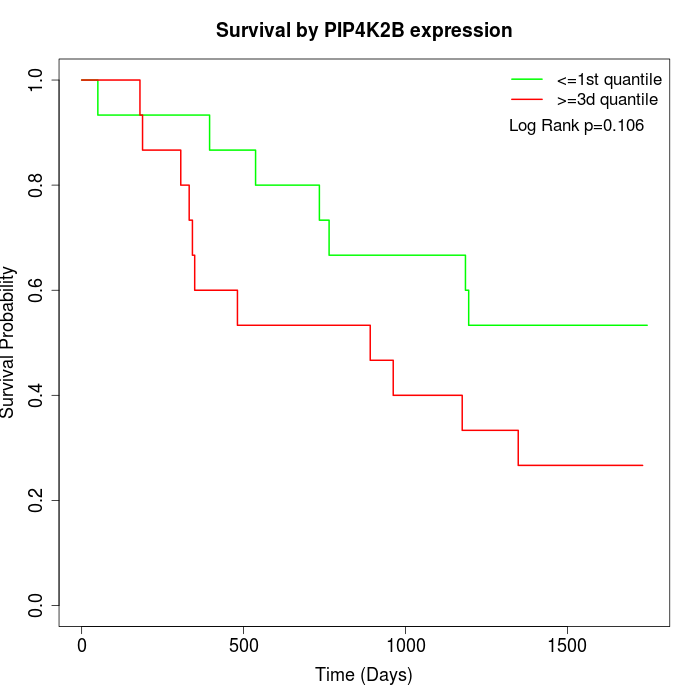
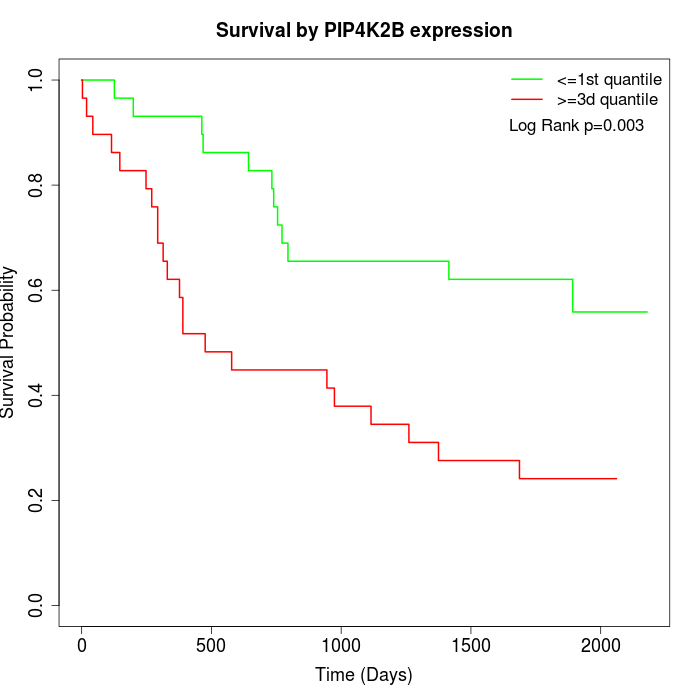
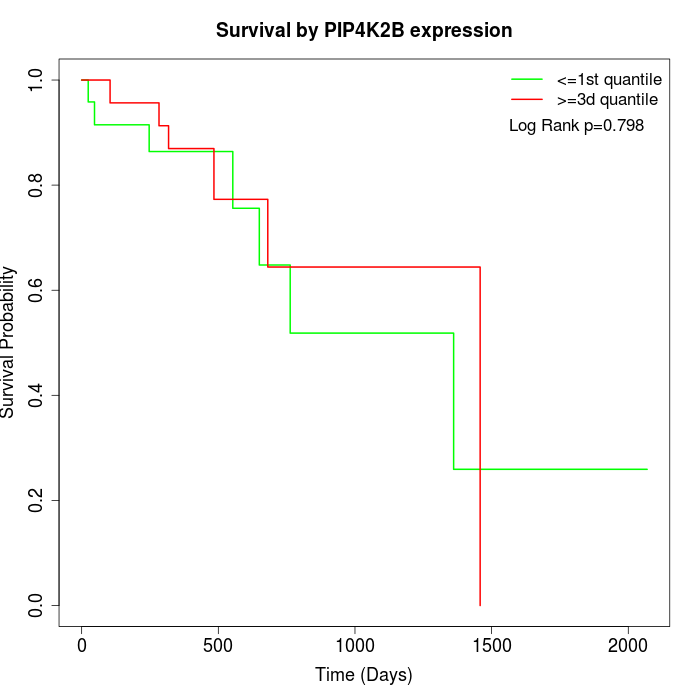
Survival by PIP4K2B expression:
Note: Click image to view full size file.
Copy number change of PIP4K2B:
| Dataset | Gene | EntrezID | Gain | Loss | Normal | Detail |
|---|---|---|---|---|---|---|
| GSE15526 | PIP4K2B | 8396 | 7 | 1 | 22 | |
| GSE20123 | PIP4K2B | 8396 | 7 | 1 | 22 | |
| GSE43470 | PIP4K2B | 8396 | 1 | 1 | 41 | |
| GSE46452 | PIP4K2B | 8396 | 34 | 0 | 25 | |
| GSE47630 | PIP4K2B | 8396 | 8 | 1 | 31 | |
| GSE54993 | PIP4K2B | 8396 | 3 | 4 | 63 | |
| GSE54994 | PIP4K2B | 8396 | 8 | 5 | 40 | |
| GSE60625 | PIP4K2B | 8396 | 4 | 0 | 7 | |
| GSE74703 | PIP4K2B | 8396 | 1 | 0 | 35 | |
| GSE74704 | PIP4K2B | 8396 | 4 | 1 | 15 | |
| TCGA | PIP4K2B | 8396 | 19 | 9 | 68 |
Total number of gains: 96; Total number of losses: 23; Total Number of normals: 369.
Somatic mutations of PIP4K2B:
Generating mutation plots.
Highly correlated genes for PIP4K2B:
Showing top 20/718 corelated genes with mean PCC>0.5.
| Gene1 | Gene2 | Mean PCC | Num. Datasets | Num. PCC<0 | Num. PCC>0.5 |
|---|---|---|---|---|---|
| PIP4K2B | TERF2 | 0.810664 | 3 | 0 | 3 |
| PIP4K2B | MLLT6 | 0.772635 | 3 | 0 | 3 |
| PIP4K2B | PTGFRN | 0.770553 | 3 | 0 | 3 |
| PIP4K2B | RBM4 | 0.755853 | 3 | 0 | 3 |
| PIP4K2B | ZZEF1 | 0.755475 | 3 | 0 | 3 |
| PIP4K2B | RHBDD2 | 0.735137 | 3 | 0 | 3 |
| PIP4K2B | PPP2R2D | 0.734991 | 3 | 0 | 3 |
| PIP4K2B | LRRK1 | 0.726915 | 3 | 0 | 3 |
| PIP4K2B | KLHL11 | 0.722132 | 3 | 0 | 3 |
| PIP4K2B | INTS3 | 0.720676 | 4 | 0 | 4 |
| PIP4K2B | MED30 | 0.719338 | 3 | 0 | 3 |
| PIP4K2B | TNRC18 | 0.718276 | 5 | 0 | 4 |
| PIP4K2B | GNAS | 0.706151 | 3 | 0 | 3 |
| PIP4K2B | CREB5 | 0.705194 | 3 | 0 | 3 |
| PIP4K2B | PTPN14 | 0.704782 | 5 | 0 | 4 |
| PIP4K2B | TNFRSF10B | 0.704772 | 3 | 0 | 3 |
| PIP4K2B | TMEM38B | 0.704766 | 3 | 0 | 3 |
| PIP4K2B | TANC2 | 0.704346 | 3 | 0 | 3 |
| PIP4K2B | RRAS2 | 0.70357 | 3 | 0 | 3 |
| PIP4K2B | ELMOD3 | 0.703167 | 3 | 0 | 3 |
For details and further investigation, click here