| Full name: vitamin D receptor | Alias Symbol: NR1I1|PPP1R163 | ||
| Type: protein-coding gene | Cytoband: 12q13.11 | ||
| Entrez ID: 7421 | HGNC ID: HGNC:12679 | Ensembl Gene: ENSG00000111424 | OMIM ID: 601769 |
| Drug and gene relationship at DGIdb | |||
VDR involved pathways:
| KEGG pathway | Description | View |
|---|---|---|
| hsa04961 | Endocrine and other factor-regulated calcium reabsorption | |
| hsa05152 | Tuberculosis |
Expression of VDR:
| Dataset | Gene | EntrezID | Probe | Log2FC | Adj.pValue | Expression |
|---|---|---|---|---|---|---|
| GSE17351 | VDR | 7421 | 204254_s_at | 0.2843 | 0.7364 | |
| GSE20347 | VDR | 7421 | 204254_s_at | -0.2345 | 0.1906 | |
| GSE23400 | VDR | 7421 | 213692_s_at | -0.1418 | 0.0012 | |
| GSE26886 | VDR | 7421 | 204254_s_at | -0.6621 | 0.0799 | |
| GSE29001 | VDR | 7421 | 204255_s_at | -0.3854 | 0.4285 | |
| GSE38129 | VDR | 7421 | 204254_s_at | 0.3292 | 0.2276 | |
| GSE45670 | VDR | 7421 | 204254_s_at | 0.3659 | 0.1507 | |
| GSE53622 | VDR | 7421 | 30975 | 0.0540 | 0.6888 | |
| GSE53624 | VDR | 7421 | 30975 | 0.0002 | 0.9984 | |
| GSE63941 | VDR | 7421 | 204254_s_at | -1.2775 | 0.0562 | |
| GSE77861 | VDR | 7421 | 204254_s_at | -0.7417 | 0.0127 | |
| GSE97050 | VDR | 7421 | A_23_P162589 | 0.8901 | 0.2864 | |
| SRP007169 | VDR | 7421 | RNAseq | -0.4488 | 0.5063 | |
| SRP008496 | VDR | 7421 | RNAseq | -0.5642 | 0.1635 | |
| SRP064894 | VDR | 7421 | RNAseq | 0.1064 | 0.5870 | |
| SRP133303 | VDR | 7421 | RNAseq | 0.3300 | 0.1064 | |
| SRP159526 | VDR | 7421 | RNAseq | -0.9457 | 0.0006 | |
| SRP193095 | VDR | 7421 | RNAseq | -0.2067 | 0.2376 | |
| SRP219564 | VDR | 7421 | RNAseq | 0.5547 | 0.4763 | |
| TCGA | VDR | 7421 | RNAseq | 0.4792 | 0.0000 |
Upregulated datasets: 0; Downregulated datasets: 0.
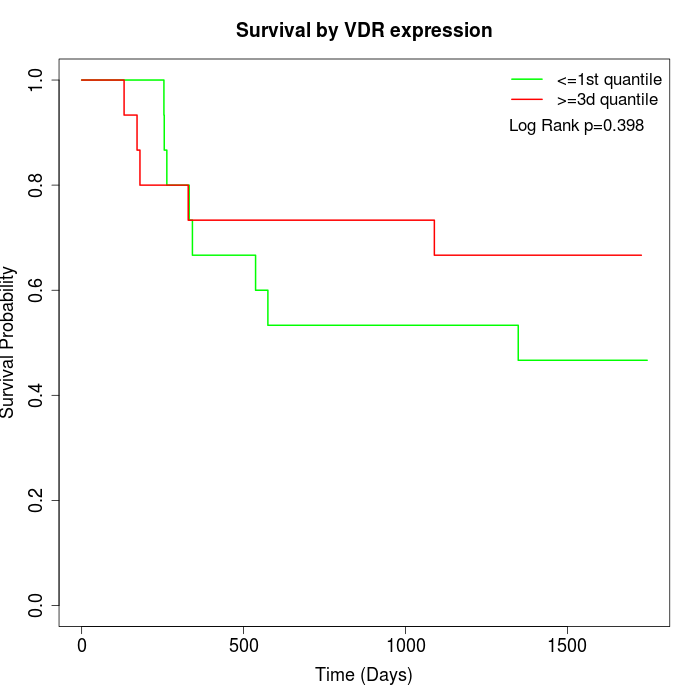
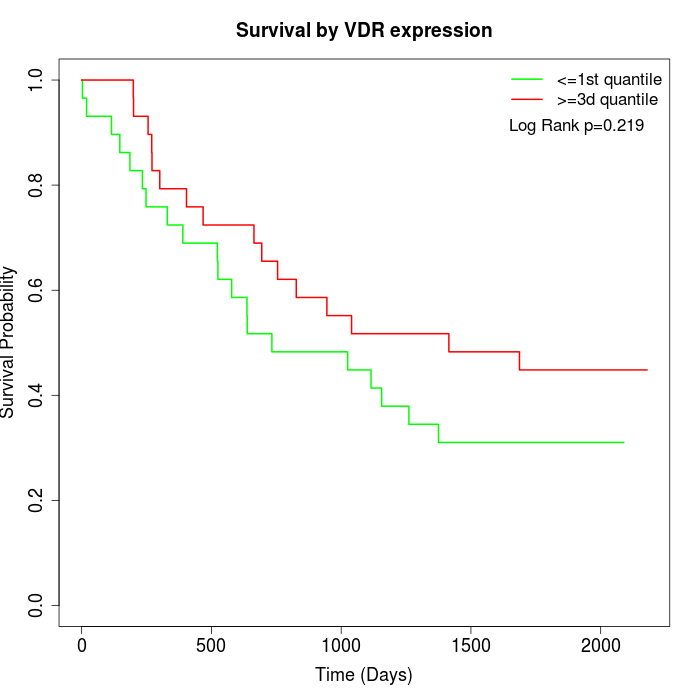
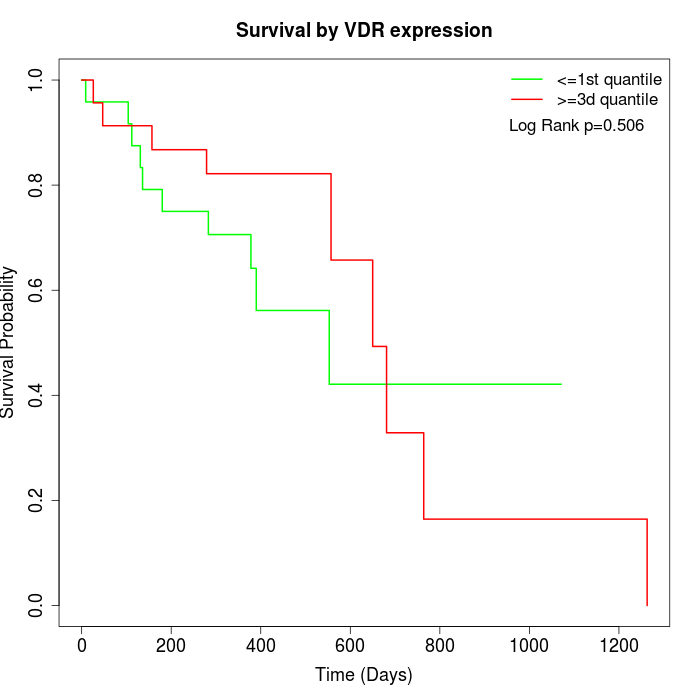
Survival by VDR expression:
Note: Click image to view full size file.
Copy number change of VDR:
| Dataset | Gene | EntrezID | Gain | Loss | Normal | Detail |
|---|---|---|---|---|---|---|
| GSE15526 | VDR | 7421 | 6 | 1 | 23 | |
| GSE20123 | VDR | 7421 | 6 | 1 | 23 | |
| GSE43470 | VDR | 7421 | 3 | 1 | 39 | |
| GSE46452 | VDR | 7421 | 8 | 1 | 50 | |
| GSE47630 | VDR | 7421 | 11 | 2 | 27 | |
| GSE54993 | VDR | 7421 | 0 | 5 | 65 | |
| GSE54994 | VDR | 7421 | 6 | 1 | 46 | |
| GSE60625 | VDR | 7421 | 0 | 0 | 11 | |
| GSE74703 | VDR | 7421 | 3 | 1 | 32 | |
| GSE74704 | VDR | 7421 | 4 | 1 | 15 | |
| TCGA | VDR | 7421 | 16 | 12 | 68 |
Total number of gains: 63; Total number of losses: 26; Total Number of normals: 399.
Somatic mutations of VDR:
Generating mutation plots.
Highly correlated genes for VDR:
Showing top 20/327 corelated genes with mean PCC>0.5.
| Gene1 | Gene2 | Mean PCC | Num. Datasets | Num. PCC<0 | Num. PCC>0.5 |
|---|---|---|---|---|---|
| VDR | STAP2 | 0.790126 | 4 | 0 | 4 |
| VDR | CAPG | 0.733341 | 8 | 0 | 7 |
| VDR | GNA15 | 0.70781 | 9 | 0 | 7 |
| VDR | FAM83C | 0.703186 | 5 | 0 | 5 |
| VDR | SH3RF2 | 0.69868 | 3 | 0 | 3 |
| VDR | PPP4R2 | 0.67337 | 3 | 0 | 3 |
| VDR | SQLE | 0.669693 | 5 | 0 | 4 |
| VDR | TMEM154 | 0.662003 | 4 | 0 | 4 |
| VDR | TACSTD2 | 0.661976 | 6 | 0 | 5 |
| VDR | APRT | 0.657609 | 4 | 0 | 3 |
| VDR | PROM2 | 0.656749 | 6 | 0 | 5 |
| VDR | ARRDC1 | 0.654371 | 5 | 0 | 4 |
| VDR | PAK6 | 0.653647 | 4 | 0 | 3 |
| VDR | S100A16 | 0.653005 | 5 | 0 | 5 |
| VDR | SGPP1 | 0.652171 | 3 | 0 | 3 |
| VDR | MAPK13 | 0.651288 | 6 | 0 | 5 |
| VDR | E2F8 | 0.649205 | 4 | 0 | 4 |
| VDR | FAM83A | 0.648553 | 7 | 0 | 7 |
| VDR | AACS | 0.648411 | 8 | 0 | 6 |
| VDR | RASSF10 | 0.648101 | 4 | 0 | 3 |
For details and further investigation, click here